HSCARG, a novel regulator of H2A ubiquitination by downregulating PRC1 ubiquitin E3 ligase activity, is essential for cell proliferation
- PMID: 24711370
- PMCID: PMC4027218
- DOI: 10.1093/nar/gku230
HSCARG, a novel regulator of H2A ubiquitination by downregulating PRC1 ubiquitin E3 ligase activity, is essential for cell proliferation
Abstract
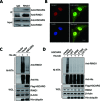
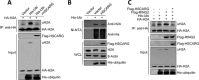
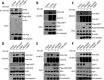
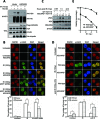
Histone H2A ubiquitination plays critical roles in transcriptional repression and deoxyribonucleic acid (DNA) damage response. More attention has been focused on ubiquitin E3 ligases of H2A, however, less is known about the negative regulators of H2A ubiquitination. Here we identified HSCARG as a new negative regulatory protein for H2A ubiquitination and found a possible link between regulator of H2A ubiquitination and cell cycle. Mechanistically, HSCARG interacts with polycomb repressive complex 1 (PRC1) and deubiquitinase USP7 and inhibits PRC1 ubiquitination in a USP7-dependent manner. As ubiquitination of PRC1 is critical for its E3 ligase activity toward H2A, HSCARG and USP7 are further shown to decrease the level of H2A ubiquitination. Moreover, we demonstrated that HSCARG is involved in DNA damage response through affecting the level of H2A ubiquitination and localization of RAP80 at lesion points. Knockout of HSCARG results in persistent activation of checkpoint signaling and leads to cell cycle arrest. This study unravels a novel mechanism for the regulation of H2A ubiquitination and elucidates how regulators of H2A ubiquitination affect cell cycle.
© The Author(s) 2014. The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

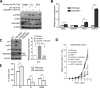
Similar articles
-
UbE2E1/UBCH6 Is a Critical in Vivo E2 for the PRC1-catalyzed Ubiquitination of H2A at Lys-119.J Biol Chem. 2017 Feb 17;292(7):2893-2902. doi: 10.1074/jbc.M116.749564. Epub 2017 Jan 10. J Biol Chem. 2017. PMID: 28073915 Free PMC article.
-
Polycomb group proteins in the DNA damage response: a link between radiation resistance and "stemness".Cell Cycle. 2011 Mar 15;10(6):883-94. doi: 10.4161/cc.10.6.14907. Epub 2011 Mar 15. Cell Cycle. 2011. PMID: 21346409
-
HSCARG downregulates NF-κB signaling by interacting with USP7 and inhibiting NEMO ubiquitination.Cell Death Dis. 2014 May 15;5(5):e1229. doi: 10.1038/cddis.2014.197. Cell Death Dis. 2014. PMID: 24832601 Free PMC article.
-
Polycomb-dependent histone H2A ubiquitination links developmental disorders with cancer.Trends Genet. 2022 Apr;38(4):333-352. doi: 10.1016/j.tig.2021.07.011. Epub 2021 Aug 20. Trends Genet. 2022. PMID: 34426021 Review.
-
Functional Link between BRCA1 and BAP1 through Histone H2A, Heterochromatin and DNA Damage Response.Curr Cancer Drug Targets. 2016;16(2):101-9. doi: 10.2174/1568009615666151030102427. Curr Cancer Drug Targets. 2016. PMID: 26517537 Review.
Cited by
-
USP7 cooperates with SCML2 to regulate the activity of PRC1.Mol Cell Biol. 2015 Apr;35(7):1157-68. doi: 10.1128/MCB.01197-14. Epub 2015 Jan 20. Mol Cell Biol. 2015. PMID: 25605328 Free PMC article.
-
Deubiquitinase OTUD6A promotes breast cancer progression by increasing TopBP1 stability and rendering tumor cells resistant to DNA-damaging therapy.Cell Death Differ. 2022 Dec;29(12):2531-2544. doi: 10.1038/s41418-022-01036-6. Epub 2022 Jun 29. Cell Death Differ. 2022. PMID: 35768646 Free PMC article.
-
Colon mucosal proteomics of ankylosing spondylitis versus gut inflammation.PLoS One. 2024 Dec 13;19(12):e0315324. doi: 10.1371/journal.pone.0315324. eCollection 2024. PLoS One. 2024. PMID: 39671362 Free PMC article.
-
Structure and functions of cellular redox sensor HSCARG/NMRAL1, a linkage among redox status, innate immunity, DNA damage response, and cancer.Free Radic Biol Med. 2020 Nov 20;160:768-774. doi: 10.1016/j.freeradbiomed.2020.09.016. Epub 2020 Sep 17. Free Radic Biol Med. 2020. PMID: 32950687 Free PMC article. Review.
-
Cellular redox sensor HSCARG negatively regulates the translesion synthesis pathway and exacerbates mammary tumorigenesis.Proc Natl Acad Sci U S A. 2019 Dec 17;116(51):25624-25633. doi: 10.1073/pnas.1910250116. Epub 2019 Dec 3. Proc Natl Acad Sci U S A. 2019. PMID: 31796584 Free PMC article.
References
-
- Goldknopf I.L., Taylor C.W., Baum R.M., Yeoman L.C., Olson M.O., Prestayko A.W., Busch H. Isolation and characterization of protein A24, a ‘histone-like’ non-histone chromosomal protein. J. Biol. Chem. 1975;250:7182–7187. - PubMed
-
- Wang H., Wang L., Erdjument-Bromage H., Vidal M., Tempst P., Jones R.S., Zhang Y. Role of histone H2A ubiquitination in Polycomb silencing. Nature. 2004;431:873–878. - PubMed
-
- Cao R., Tsukada Y., Zhang Y. Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol. Cell. 2005;20:845–854. - PubMed
-
- Mailand N., Bekker-Jensen S., Faustrup H., Melander F., Bartek J., Lukas C., Lukas J. RNF8 ubiquitylates histones at DNA double-strand breaks and promotes assembly of repair proteins. Cell. 2007;131:887–900. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

