A novel probe for phosphatidylinositol 4-phosphate reveals multiple pools beyond the Golgi
- PMID: 24711504
- PMCID: PMC3987136
- DOI: 10.1083/jcb.201312072
A novel probe for phosphatidylinositol 4-phosphate reveals multiple pools beyond the Golgi
Abstract
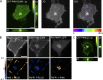
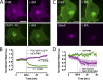
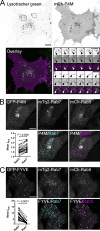
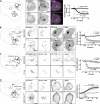
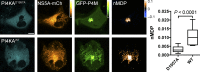
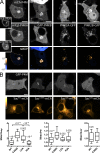
Polyphosphoinositides are an important class of lipid that recruit specific effector proteins to organelle membranes. One member, phosphatidylinositol 4-phosphate (PtdIns4P) has been localized to Golgi membranes based on the distribution of lipid binding modules from PtdIns4P effector proteins. However, these probes may be biased by additional interactions with other Golgi-specific determinants. In this paper, we derive a new PtdIns4P biosensor using the PtdIns4P binding of SidM (P4M) domain of the secreted effector protein SidM from the bacterial pathogen Legionella pneumophila. PtdIns4P was necessary and sufficient for localization of P4M, which revealed pools of the lipid associated not only with the Golgi but also with the plasma membrane and Rab7-positive late endosomes/lysosomes. PtdIns4P distribution was determined by the localization and activities of both its anabolic and catabolic enzymes. Therefore, P4M reports a wider cellular distribution of PtdIns4P than previous probes and therefore will be valuable for dissecting the biological functions of PtdIns4P in its assorted membrane compartments.
Figures


Similar articles
-
Rab1 guanine nucleotide exchange factor SidM is a major phosphatidylinositol 4-phosphate-binding effector protein of Legionella pneumophila.J Biol Chem. 2009 Feb 20;284(8):4846-56. doi: 10.1074/jbc.M807505200. Epub 2008 Dec 17. J Biol Chem. 2009. PMID: 19095644 Free PMC article.
-
Multiphasic dynamics of phosphatidylinositol 4-phosphate during phagocytosis.Mol Biol Cell. 2017 Jan 1;28(1):128-140. doi: 10.1091/mbc.E16-06-0451. Epub 2016 Nov 9. Mol Biol Cell. 2017. PMID: 28035045 Free PMC article.
-
VAMP associated proteins are required for autophagic and lysosomal degradation by promoting a PtdIns4P-mediated endosomal pathway.Autophagy. 2019 Jul;15(7):1214-1233. doi: 10.1080/15548627.2019.1580103. Epub 2019 Feb 20. Autophagy. 2019. PMID: 30741620 Free PMC article.
-
Phosphoinositides in Golgi complex function.Subcell Biochem. 2012;59:255-70. doi: 10.1007/978-94-007-3015-1_8. Subcell Biochem. 2012. PMID: 22374093 Review.
-
Coordination of Golgi functions by phosphatidylinositol 4-kinases.Trends Cell Biol. 2011 Feb;21(2):113-21. doi: 10.1016/j.tcb.2010.10.002. Epub 2010 Nov 4. Trends Cell Biol. 2011. PMID: 21282087 Free PMC article. Review.
Cited by
-
Lipid composition of the cancer cell membrane.J Bioenerg Biomembr. 2020 Oct;52(5):321-342. doi: 10.1007/s10863-020-09846-4. Epub 2020 Jul 26. J Bioenerg Biomembr. 2020. PMID: 32715369 Free PMC article. Review.
-
β-arrestin-dependent PI(4,5)P2 synthesis boosts GPCR endocytosis.Proc Natl Acad Sci U S A. 2021 Apr 27;118(17):e2011023118. doi: 10.1073/pnas.2011023118. Proc Natl Acad Sci U S A. 2021. PMID: 33879605 Free PMC article.
-
Cholesterol transfer via endoplasmic reticulum contacts mediates lysosome damage repair.EMBO J. 2022 Dec 15;41(24):e112677. doi: 10.15252/embj.2022112677. Epub 2022 Nov 21. EMBO J. 2022. PMID: 36408828 Free PMC article.
-
PI(4,5)P2 diffuses freely in the plasma membrane even within high-density effector protein complexes.J Cell Biol. 2023 Feb 6;222(2):e202204099. doi: 10.1083/jcb.202204099. Epub 2022 Nov 23. J Cell Biol. 2023. PMID: 36416724 Free PMC article.
-
OSBP is a major determinant of Golgi phosphatidylinositol 4-phosphate homeostasis.bioRxiv [Preprint]. 2024 Jan 25:2023.12.21.572879. doi: 10.1101/2023.12.21.572879. bioRxiv. 2024. Update in: Contact (Thousand Oaks). 2024 Feb 22;7:25152564241232196. doi: 10.1177/25152564241232196. PMID: 38187665 Free PMC article. Updated. Preprint.
References
-
- Balla A., Tuymetova G., Tsiomenko A., Várnai P., Balla T. 2005. A plasma membrane pool of phosphatidylinositol 4-phosphate is generated by phosphatidylinositol 4-kinase type-III alpha: studies with the PH domains of the oxysterol binding protein and FAPP1. Mol. Biol. Cell. 16:1282–1295 10.1091/mbc.E04-07-0578 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

