Streptococcus pneumoniae detects and responds to foreign bacterial peptide fragments in its environment
- PMID: 24718598
- PMCID: PMC4043112
- DOI: 10.1098/rsob.130224
Streptococcus pneumoniae detects and responds to foreign bacterial peptide fragments in its environment
Abstract
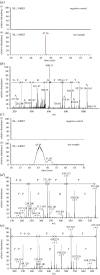
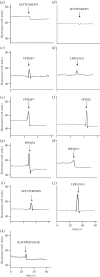
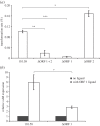
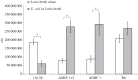
Streptococcus pneumoniae is an important cause of bacterial meningitis and pneumonia but usually colonizes the human nasopharynx harmlessly. As this niche is simultaneously populated by other bacterial species, we looked for a role and pathway of communication between pneumococci and other species. This paper shows that two proteins of non-encapsulated S. pneumoniae, AliB-like ORF 1 and ORF 2, bind specifically to peptides matching other species resulting in changes in the pneumococci. AliB-like ORF 1 binds specifically peptide SETTFGRDFN, matching 50S ribosomal subunit protein L4 of Enterobacteriaceae, and facilitates upregulation of competence for genetic transformation. AliB-like ORF 2 binds specifically peptides containing sequence FPPQS, matching proteins of Prevotella species common in healthy human nasopharyngeal microbiota. We found that AliB-like ORF 2 mediates the early phase of nasopharyngeal colonization in vivo. The ability of S. pneumoniae to bind and respond to peptides of other bacterial species occupying the same host niche may play a key role in adaptation to its environment and in interspecies communication. These findings reveal a completely new concept of pneumococcal interspecies communication which may have implications for communication between other bacterial species and for future interventional therapeutics.
Keywords: Streptococcus pneumoniae, bacteria; interspecies communication; non-encapsulated; peptide.
Figures
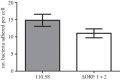

References
-
- Ryan R, Dow J. 2008. Diffusible signals and interspecies communication in bacteria. Microbiology 154, 1845–1858. (doi:10.1099/mic.0.2008/017871-0) - DOI - PubMed
-
- Schauder S, Bassler B. 2001. The languages of bacteria. Genes Dev. 15, 1468–1480. (doi:10.1101/gad.899601) - DOI - PubMed
-
- Lam H, Oh D-C, Cava F, Takacs C, Clardy J, Pedro Md, Waldor M. 2009. d-Amino acids govern stationary phase cell wall remodeling in bacteria. Science 325, 1552–1555. (doi:10.1126/science.1178123) - DOI - PMC - PubMed
-
- Kolodkin-Gal I, Romero D, Cao S, Clardy J, Kolter R, Losick R. 2010. d-Amino acids trigger biofilm disassembly. Science 328, 627–629. (doi:10.1126/science.1188628) - DOI - PMC - PubMed
-
- Cava F, Pedro Md, Lam H, Davis B, Waldor M. 2011. Distinct pathways for modification of the bacterial cell wall by non-canonical d-amino acids. The EMBO J. 30, 3442–3453. (doi:10.1038/emboj.2011.246) - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
