Only minimal regions of tomato yellow leaf curl virus (TYLCV) are required for replication, expression and movement
- PMID: 24719195
- PMCID: PMC4147252
- DOI: 10.1007/s00705-014-2066-7
Only minimal regions of tomato yellow leaf curl virus (TYLCV) are required for replication, expression and movement
Abstract
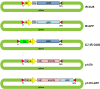
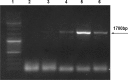
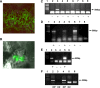
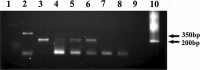
The IL-60 platform, consisting of a disarmed form of tomato yellow leaf curl virus (TYLCV) and auxiliary components, was previously developed as a nontransgenic universal vector system for gene expression and silencing that can express an entire operon in plants. IL-60 does not allow rolling-circle replication; hence, production of viral single-stranded (ss) DNA progeny is prevented. We used this double-stranded (ds) DNA-restricted platform (uncoupled from the dsDNA→ssDNA replication phase of progeny viral DNA) for functional genomics studies of TYLCV. We report that the noncoding 314-bp intergenic region (IR) is the only viral element required for viral dsDNA replication. None of the viral genes are required, suggesting recruitment of host factors that recognize the IR. We further show that IR-carrying reporter genes are also capable of replication but remain confined to the cells into which they were introduced. Only two sense-oriented viral genes (V1 and V2) need to be added to the IR-carrying construct for expression and movement. Hence, any IR-dsDNA construct supplemented with V1 and V2 becomes a replication-competent, mobile and expressing plant plasmid. All viral functions (replication, expression and movement) are determined by the IR and the sense-oriented genes. The complementary-oriented viral genes have auxiliary roles in the late phase of the virus "life cycle". The previously reported involvement of some viral genes in expression and movement is therefore revised.
Figures


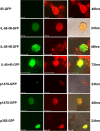

Similar articles
-
Tomato yellow leaf curl virus intergenic siRNAs target a host long noncoding RNA to modulate disease symptoms.PLoS Pathog. 2019 Jan 22;15(1):e1007534. doi: 10.1371/journal.ppat.1007534. eCollection 2019 Jan. PLoS Pathog. 2019. PMID: 30668603 Free PMC article.
-
TYLCV-Is movement in planta does not require V2 protein.Virology. 2015 Mar;477:56-60. doi: 10.1016/j.virol.2015.01.007. Epub 2015 Feb 5. Virology. 2015. PMID: 25644513
-
A peptide derived from enzymatic digestion of globulins from amaranth shows strong affinity binding to the replication origin of Tomato yellow leaf curl virus reducing viral replication in Nicotiana benthamiana.Pestic Biochem Physiol. 2018 Feb;145:56-65. doi: 10.1016/j.pestbp.2018.01.005. Epub 2018 Feb 4. Pestic Biochem Physiol. 2018. PMID: 29482732
-
Replication of Tomato Yellow Leaf Curl Virus in Its Whitefly Vector, Bemisia tabaci.J Virol. 2015 Oct;89(19):9791-803. doi: 10.1128/JVI.00779-15. Epub 2015 Jul 15. J Virol. 2015. PMID: 26178995 Free PMC article.
-
Discovering host genes involved in the infection by the Tomato Yellow Leaf Curl Virus complex and in the establishment of resistance to the virus using Tobacco Rattle Virus-based post transcriptional gene silencing.Viruses. 2013 Mar 22;5(3):998-1022. doi: 10.3390/v5030998. Viruses. 2013. PMID: 23524390 Free PMC article. Review.
Cited by
-
Regulation of Flowering Timing by ABA-NnSnRK1 Signaling Pathway in Lotus.Int J Mol Sci. 2021 Apr 10;22(8):3932. doi: 10.3390/ijms22083932. Int J Mol Sci. 2021. PMID: 33920313 Free PMC article.
-
New biotechnological tools to accelerate scab-resistance trait transfer to apple.Genet Mol Biol. 2017;40(1 suppl 1):305-311. doi: 10.1590/1678-4685-GMB-2016-0043. Epub 2017 Feb 13. Genet Mol Biol. 2017. PMID: 28199444 Free PMC article.
-
Plant biomacromolecule delivery methods in the 21st century.Front Genome Ed. 2022 Oct 14;4:1011934. doi: 10.3389/fgeed.2022.1011934. eCollection 2022. Front Genome Ed. 2022. PMID: 36311974 Free PMC article. Review.
-
CRISPR/Cas systems versus plant viruses: engineering plant immunity and beyond.Plant Physiol. 2021 Aug 3;186(4):1770-1785. doi: 10.1093/plphys/kiab220. Plant Physiol. 2021. PMID: 35237805 Free PMC article.
-
CRISPR/Cas9-mediated viral interference in plants.Genome Biol. 2015 Nov 11;16:238. doi: 10.1186/s13059-015-0799-6. Genome Biol. 2015. PMID: 26556628 Free PMC article.
References
-
- Stanley J, Bisaro DM, Briddon RW, Brown JK, Faucuet CM, Harrison BD, et al. Family geminiviridae. In: Faucuet CM, Maniloff J, Desselberger U, Ball LA, et al., editors. Virus taxonomy. English report of the International Committee on Taxonomy of Viruses. San Diego: Elsevier Academic Press; 2005. pp. 301–326.
-
- Jeske H. Geminiviruses. Curr Top Microbiol Immunol. 2009;331:185–226. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

