Template-based structure modeling of protein-protein interactions
- PMID: 24721449
- PMCID: PMC3984454
- DOI: 10.1016/j.sbi.2013.11.005
Template-based structure modeling of protein-protein interactions
Abstract
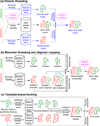
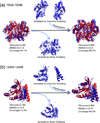
The structure of protein-protein complexes can be constructed by using the known structure of other protein complexes as a template. The complex structure templates are generally detected either by homology-based sequence alignments or, given the structure of monomer components, by structure-based comparisons. Critical improvements have been made in recent years by utilizing interface recognition and by recombining monomer and complex template libraries. Encouraging progress has also been witnessed in genome-wide applications of template-based modeling, with modeling accuracy comparable to high-throughput experimental data. Nevertheless, bottlenecks exist due to the incompleteness of the protein-protein complex structure library and the lack of methods for distant homologous template identification and full-length complex structure refinement.
Copyright © 2013 Elsevier Ltd. All rights reserved.
Figures

References
-
- Uetz P, Giot L, Cagney G, Mansfield TA, Judson RS, Knight JR, Lockshon D, Narayan V, Srinivasan M, Pochart P, et al. A comprehensive analysis of protein-protein interactions in Saccharomyces cerevisiae. Nature. 2000;403:623–627. - PubMed
-
- Rain JC, Selig L, De Reuse H, Battaglia V, Reverdy C, Simon S, Lenzen G, Petel F, Wojcik J, Schachter V, et al. The protein-protein interaction map of Helicobacter pylori. Nature. 2001;409:211–215. - PubMed
-
- Mosca R, Pons T, Ceol A, Valencia A, Aloy P. Towards a detailed atlas of protein-protein interactions. Curr Opin Struct Biol. 2013 - PubMed
-
- Stein A, Mosca R, Aloy P. Three-dimensional modeling of protein interactions and complexes is going 'omics. Curr Opin Struct Biol. 2011;21:200–208. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

