Ca2+ binding enhanced mechanical stability of an archaeal crystallin
- PMID: 24728085
- PMCID: PMC3984160
- DOI: 10.1371/journal.pone.0094513
Ca2+ binding enhanced mechanical stability of an archaeal crystallin
Erratum in
- PLoS One. 2014;9(7):e102924
Abstract
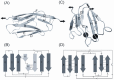
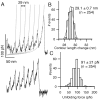
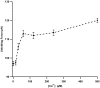
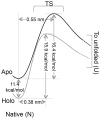
Structural topology plays an important role in protein mechanical stability. Proteins with β-sandwich topology consisting of Greek key structural motifs, for example, I27 of muscle titin and (10)FNIII of fibronectin, are mechanically resistant as shown by single-molecule force spectroscopy (SMFS). In proteins with β-sandwich topology, if the terminal strands are directly connected by backbone H-bonding then this geometry can serve as a "mechanical clamp". Proteins with this geometry are shown to have very high unfolding forces. Here, we set out to explore the mechanical properties of a protein, M-crystallin, which belongs to β-sandwich topology consisting of Greek key motifs but its overall structure lacks the "mechanical clamp" geometry at the termini. M-crystallin is a Ca(2+) binding protein from Methanosarcina acetivorans that is evolutionarily related to the vertebrate eye lens β and γ-crystallins. We constructed an octamer of crystallin, (M-crystallin)8, and using SMFS, we show that M-crystallin unfolds in a two-state manner with an unfolding force ∼ 90 pN (at a pulling speed of 1000 nm/sec), which is much lower than that of I27. Our study highlights that the β-sandwich topology proteins with a different strand-connectivity than that of I27 and (10)FNIII, as well as lacking "mechanical clamp" geometry, can be mechanically resistant. Furthermore, Ca(2+) binding not only stabilizes M-crystallin by 11.4 kcal/mol but also increases its unfolding force by ∼ 35 pN at the same pulling speed. The differences in the mechanical properties of apo and holo M-crystallins are further characterized using pulling speed dependent measurements and they show that Ca(2+) binding reduces the unfolding potential width from 0.55 nm to 0.38 nm. These results are explained using a simple two-state unfolding energy landscape.
Conflict of interest statement
Figures


Similar articles
-
Solution structure and calcium-binding properties of M-crystallin, a primordial betagamma-crystallin from archaea.J Mol Biol. 2009 Feb 27;386(3):675-89. doi: 10.1016/j.jmb.2008.12.058. Epub 2008 Dec 31. J Mol Biol. 2009. PMID: 19138688
-
Iterative cloning, overexpression, purification and isotopic labeling of an engineered dimer of a Ca(2+)-binding protein of the βγ-crystallin superfamily from Methanosarcina acetivorans.Protein Expr Purif. 2012 Jul;84(1):116-22. doi: 10.1016/j.pep.2012.04.024. Epub 2012 May 9. Protein Expr Purif. 2012. PMID: 22579642
-
Calcium binding properties of gamma-crystallin: calcium ion binds at the Greek key beta gamma-crystallin fold.J Biol Chem. 2001 Oct 19;276(42):38464-71. doi: 10.1074/jbc.M102164200. Epub 2001 Aug 13. J Biol Chem. 2001. PMID: 11502736
-
Ca2+-binding motif of βγ-crystallins.J Biol Chem. 2014 Apr 18;289(16):10958-10966. doi: 10.1074/jbc.O113.539569. Epub 2014 Feb 24. J Biol Chem. 2014. PMID: 24567326 Free PMC article. Review.
-
Function and Aggregation in Structural Eye Lens Crystallins.Acc Chem Res. 2020 Apr 21;53(4):863-874. doi: 10.1021/acs.accounts.0c00014. Epub 2020 Apr 9. Acc Chem Res. 2020. PMID: 32271004 Free PMC article. Review.
Cited by
-
Ligand-induced changes of the apparent transition-state position in mechanical protein unfolding.Biophys J. 2015 Jul 21;109(2):365-72. doi: 10.1016/j.bpj.2015.06.009. Biophys J. 2015. PMID: 26200872 Free PMC article.
-
Single-molecule Force Spectroscopy Predicts a Misfolded, Domain-swapped Conformation in human γD-Crystallin Protein.J Biol Chem. 2016 Feb 19;291(8):4226-35. doi: 10.1074/jbc.M115.673871. Epub 2015 Dec 24. J Biol Chem. 2016. PMID: 26703476 Free PMC article.
-
Differences in the mechanical unfolding pathways of apo- and copper-bound azurins.Sci Rep. 2018 Jan 31;8(1):1989. doi: 10.1038/s41598-018-19755-7. Sci Rep. 2018. PMID: 29386517 Free PMC article.
-
Cation-induced shape programming and morphing in protein-based hydrogels.Sci Adv. 2020 Apr 29;6(18):eaba6112. doi: 10.1126/sciadv.aba6112. eCollection 2020 May. Sci Adv. 2020. PMID: 32494690 Free PMC article.
-
Unusually high mechanical stability of bacterial adhesin extender domains having calcium clamps.PLoS One. 2017 Apr 4;12(4):e0174682. doi: 10.1371/journal.pone.0174682. eCollection 2017. PLoS One. 2017. PMID: 28376122 Free PMC article.
References
-
- Oberhauser AF, Badilla-Fernandez C, Carrion-Vazquez M, Fernandez JM (2002) The mechanical hierarchies of fibronectin observed with single-molecule AFM. J Mol Biol 319: 433–447. - PubMed
-
- Li H, Fernandez JM (2003) Mechanical design of the first proximal Ig domain of human cardiac titin revealed by single molecule force spectroscopy. J Mol Biol 334: 75–86. - PubMed
-
- Li L, Huang HH, Badilla CL, Fernandez JM (2005) Mechanical unfolding intermediates observed by single-molecule force spectroscopy in a fibronectin type III module. J Mol Biol 345: 817–826. - PubMed
-
- Ng SP, Rounsevell RW, Steward A, Geierhaas CD, Williams PM, et al. (2005) Mechanical unfolding of TNfn3: the unfolding pathway of a fnIII domain probed by protein engineering, AFM and MD simulation. J Mol Biol 350: 776–789. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

