Polycomb proteins control proliferation and transformation independently of cell cycle checkpoints by regulating DNA replication
- PMID: 24728135
- PMCID: PMC3996544
- DOI: 10.1038/ncomms4649
Polycomb proteins control proliferation and transformation independently of cell cycle checkpoints by regulating DNA replication
Abstract
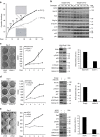
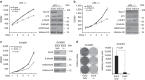
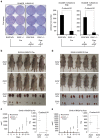
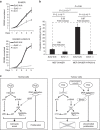
The ability of PRC1 and PRC2 to promote proliferation is a main feature that links polycomb (PcG) activity to cancer. PcGs silence the expression of the tumour suppressor locus Ink4a/Arf, whose products positively regulate pRb and p53 functions. Enhanced PcG activity is a frequent feature of human tumours, and PcG inhibition has been proposed as a strategy for cancer treatment. However, the recurrent inactivation of pRb/p53 responses in human cancers raises a question regarding the ability of PcG proteins to affect cellular proliferation independently from this checkpoint. Here we demonstrate that PRCs regulate cellular proliferation and transformation independently of the Ink4a/Arf-pRb-p53 pathway. We provide evidence that PRCs localize at replication forks, and that loss of their function directly affects the progression and symmetry of DNA replication forks. Thus, we have identified a novel activity by which PcGs can regulate cell proliferation independently of major cell cycle restriction checkpoints.
Conflict of interest statement
K.H. is a cofounder of EpiTherapeutics and has shares and warrants in the company. The remaining authors declare no competing financial interests.
Figures




References
-
- Hanahan D. & Weinberg R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011). - PubMed
-
- Medema R. H. & Macurek L. Checkpoint control and cancer. Oncogene 31, 2601–2613 (2012). - PubMed
-
- Hahn W. C. et al. Creation of human tumour cells with defined genetic elements. Nature 400, 464–468 (1999). - PubMed
-
- Ventura A. et al. Restoration of p53 function leads to tumour regression in vivo. Nature 445, 661–665 (2007). - PubMed
-
- Serrano M. et al. Role of the INK4a locus in tumour suppression and cell mortality. Cell 85, 27–37 (1996). - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

