The USDA barley core collection: genetic diversity, population structure, and potential for genome-wide association studies
- PMID: 24732668
- PMCID: PMC3986206
- DOI: 10.1371/journal.pone.0094688
The USDA barley core collection: genetic diversity, population structure, and potential for genome-wide association studies
Abstract
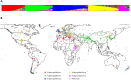
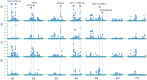
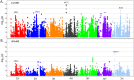
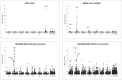
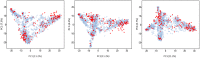
New sources of genetic diversity must be incorporated into plant breeding programs if they are to continue increasing grain yield and quality, and tolerance to abiotic and biotic stresses. Germplasm collections provide a source of genetic and phenotypic diversity, but characterization of these resources is required to increase their utility for breeding programs. We used a barley SNP iSelect platform with 7,842 SNPs to genotype 2,417 barley accessions sampled from the USDA National Small Grains Collection of 33,176 accessions. Most of the accessions in this core collection are categorized as landraces or cultivars/breeding lines and were obtained from more than 100 countries. Both STRUCTURE and principal component analysis identified five major subpopulations within the core collection, mainly differentiated by geographical origin and spike row number (an inflorescence architecture trait). Different patterns of linkage disequilibrium (LD) were found across the barley genome and many regions of high LD contained traits involved in domestication and breeding selection. The genotype data were used to define 'mini-core' sets of accessions capturing the majority of the allelic diversity present in the core collection. These 'mini-core' sets can be used for evaluating traits that are difficult or expensive to score. Genome-wide association studies (GWAS) of 'hull cover', 'spike row number', and 'heading date' demonstrate the utility of the core collection for locating genetic factors determining important phenotypes. The GWAS results were referenced to a new barley consensus map containing 5,665 SNPs. Our results demonstrate that GWAS and high-density SNP genotyping are effective tools for plant breeders interested in accessing genetic diversity in large germplasm collections.
Conflict of interest statement
Figures
References
-
- Badr A, Müller K, Schäfer-Pregl R, El Rabey H, Effgen S, et al. (2000) On the origin and domestication history of Barley (Hordeum vulgare). Mol Biol Evol 17: 499–510. - PubMed
-
- Baik B-K, Ullrich SE (2008) Barley for food: characteristics, improvement, and renewed interest. J Cereal Sci 48: 233–242.
-
- AbuMweis SS, Jew S, Ames NP (2010) β-glucan from barley and its lipid-lowering capacity: a meta-analysis of randomized, controlled trials. Eur J Clin Nutr 64: 1472–1480. - PubMed
-
- Brockman DA, Chen X, Gallaher DD (2013) Consumption of a high β-glucan barley flour improves glucose control and fatty liver and increases muscle acylcarnitines in the Zucker diabetic fatty rat. Eur J Clin Nutr 52: 1743–1753. - PubMed
-
- Sullivan P, Arendt E, Gallagher E (2013) The increasing use of barley and barley by-products in the production of healthier baked goods. Trends Food Sci Technol 29: 124–134.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

