Arabidopsis Class I α-Mannosidases MNS4 and MNS5 Are Involved in Endoplasmic Reticulum-Associated Degradation of Misfolded Glycoproteins
- PMID: 24737672
- PMCID: PMC4036581
- DOI: 10.1105/tpc.114.123216
Arabidopsis Class I α-Mannosidases MNS4 and MNS5 Are Involved in Endoplasmic Reticulum-Associated Degradation of Misfolded Glycoproteins
Abstract
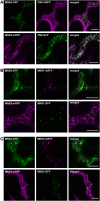
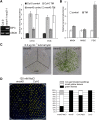
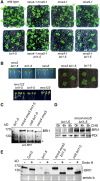
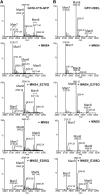
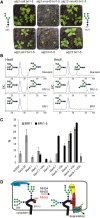
To ensure that aberrantly folded proteins are cleared from the endoplasmic reticulum (ER), all eukaryotic cells possess a mechanism known as endoplasmic reticulum-associated degradation (ERAD). Many secretory proteins are N-glycosylated, and despite some recent progress, little is known about the mechanism that selects misfolded glycoproteins for degradation in plants. Here, we investigated the role of Arabidopsis thaliana class I α-mannosidases (MNS1 to MNS5) in glycan-dependent ERAD. Our genetic and biochemical data show that the two ER-resident proteins MNS4 and MNS5 are involved in the degradation of misfolded variants of the heavily glycosylated brassinosteroid receptor, BRASSINOSTEROID INSENSITIVE1, while MNS1 to MNS3 appear dispensable for this ERAD process. By contrast, N-glycan analysis of different mns mutant combinations revealed that MNS4 and MNS5 are not involved in regular N-glycan processing of properly folded secretory glycoproteins. Overexpression of MNS4 or MNS5 together with ER-retained glycoproteins indicates further that both enzymes can convert Glc0-1Man8-9GlcNAc2 into N-glycans with a terminal α1,6-linked Man residue in the C-branch. Thus, MNS4 and MNS5 function in the formation of unique N-glycan structures that are specifically recognized by other components of the ERAD machinery, which ultimately results in the disposal of misfolded glycoproteins.
© 2014 American Society of Plant Biologists. All rights reserved.
Figures




Similar articles
-
A Non-redundant Function of MNS5: A Class I α-1, 2 Mannosidase, in the Regulation of Endoplasmic Reticulum-Associated Degradation of Misfolded Glycoproteins.Front Plant Sci. 2022 Apr 19;13:873688. doi: 10.3389/fpls.2022.873688. eCollection 2022. Front Plant Sci. 2022. PMID: 35519817 Free PMC article.
-
A context-independent N-glycan signal targets the misfolded extracellular domain of Arabidopsis STRUBBELIG to endoplasmic-reticulum-associated degradation.Biochem J. 2014 Dec 15;464(3):401-11. doi: 10.1042/BJ20141057. Biochem J. 2014. PMID: 25251695 Free PMC article.
-
Class I alpha-mannosidases are required for N-glycan processing and root development in Arabidopsis thaliana.Plant Cell. 2009 Dec;21(12):3850-67. doi: 10.1105/tpc.109.072363. Epub 2009 Dec 18. Plant Cell. 2009. PMID: 20023195 Free PMC article.
-
The Crucial Role of Demannosylating Asparagine-Linked Glycans in ERADicating Misfolded Glycoproteins in the Endoplasmic Reticulum.Front Plant Sci. 2021 Jan 12;11:625033. doi: 10.3389/fpls.2020.625033. eCollection 2020. Front Plant Sci. 2021. PMID: 33510762 Free PMC article. Review.
-
Glycan regulation of ER-associated degradation through compartmentalization.Semin Cell Dev Biol. 2015 May;41:99-109. doi: 10.1016/j.semcdb.2014.11.006. Epub 2014 Nov 24. Semin Cell Dev Biol. 2015. PMID: 25460542 Review.
Cited by
-
Overexpression of wheat α-mannosidase gene TaMP impairs salt tolerance in transgenic Brachypodium distachyon.Plant Cell Rep. 2020 May;39(5):653-667. doi: 10.1007/s00299-020-02522-2. Epub 2020 Mar 2. Plant Cell Rep. 2020. PMID: 32123996
-
Plant Virus Infection and the Ubiquitin Proteasome Machinery: Arms Race along the Endoplasmic Reticulum.Viruses. 2016 Nov 19;8(11):314. doi: 10.3390/v8110314. Viruses. 2016. PMID: 27869775 Free PMC article. Review.
-
Biological significance of complex N-glycans in plants and their impact on plant physiology.Front Plant Sci. 2014 Jul 22;5:363. doi: 10.3389/fpls.2014.00363. eCollection 2014. Front Plant Sci. 2014. PMID: 25101107 Free PMC article. Review.
-
Trafficking and localization of Golgi-resident N-glycan processing enzymes in plants.Front Plant Sci. 2025 Jul 25;16:1624949. doi: 10.3389/fpls.2025.1624949. eCollection 2025. Front Plant Sci. 2025. PMID: 40786940 Free PMC article. Review.
-
An endoplasmic reticulum-associated degradation-related E2-E3 enzyme pair controls grain size and weight through the brassinosteroid signaling pathway in rice.Plant Cell. 2023 Mar 15;35(3):1076-1091. doi: 10.1093/plcell/koac364. Plant Cell. 2023. PMID: 36519262 Free PMC article.
References
-
- Aebi M., Bernasconi R., Clerc S., Molinari M. (2010). N-Glycan structures: Recognition and processing in the ER. Trends Biochem. Sci. 35: 74–82 - PubMed
-
- Brandizzi F., Hanton S., DaSilva L.L., Boevink P., Evans D., Oparka K., Denecke J., Hawes C. (2003). ER quality control can lead to retrograde transport from the ER lumen to the cytosol and the nucleoplasm in plants. Plant J. 34: 269–281 - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

