A general approach for haplotype phasing across the full spectrum of relatedness
- PMID: 24743097
- PMCID: PMC3990520
- DOI: 10.1371/journal.pgen.1004234
A general approach for haplotype phasing across the full spectrum of relatedness
Abstract
Many existing cohorts contain a range of relatedness between genotyped individuals, either by design or by chance. Haplotype estimation in such cohorts is a central step in many downstream analyses. Using genotypes from six cohorts from isolated populations and two cohorts from non-isolated populations, we have investigated the performance of different phasing methods designed for nominally 'unrelated' individuals. We find that SHAPEIT2 produces much lower switch error rates in all cohorts compared to other methods, including those designed specifically for isolated populations. In particular, when large amounts of IBD sharing is present, SHAPEIT2 infers close to perfect haplotypes. Based on these results we have developed a general strategy for phasing cohorts with any level of implicit or explicit relatedness between individuals. First SHAPEIT2 is run ignoring all explicit family information. We then apply a novel HMM method (duoHMM) to combine the SHAPEIT2 haplotypes with any family information to infer the inheritance pattern of each meiosis at all sites across each chromosome. This allows the correction of switch errors, detection of recombination events and genotyping errors. We show that the method detects numbers of recombination events that align very well with expectations based on genetic maps, and that it infers far fewer spurious recombination events than Merlin. The method can also detect genotyping errors and infer recombination events in otherwise uninformative families, such as trios and duos. The detected recombination events can be used in association scans for recombination phenotypes. The method provides a simple and unified approach to haplotype estimation, that will be of interest to researchers in the fields of human, animal and plant genetics.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
 ,
,  ,
,  and
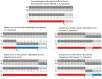
and  denotes the two parental and child haplotypes and
denotes the two parental and child haplotypes and  denotes the pattern of gene flow. Top: Correctly inferred haplotypes in a region of a true recombination event that causes a
denotes the pattern of gene flow. Top: Correctly inferred haplotypes in a region of a true recombination event that causes a  transition in the duo HMM. The other 4 examples in the figure add SEs to these true parental and child haplotypes. Middle left: addition of a SE in the child's haplotypes that causes a
transition in the duo HMM. The other 4 examples in the figure add SEs to these true parental and child haplotypes. Middle left: addition of a SE in the child's haplotypes that causes a  transition. Middle right: addition of a SE in the parent's haplotypes that causes a
transition. Middle right: addition of a SE in the parent's haplotypes that causes a  transition. Bottom left: addition of a SE in the parent's haplotypes at the site of the recombination event that causes the
transition. Bottom left: addition of a SE in the parent's haplotypes at the site of the recombination event that causes the  transition to be missed. Bottom right: addition of a SE in both the child's and parent's haplotypes at the same position that causes a
transition to be missed. Bottom right: addition of a SE in both the child's and parent's haplotypes at the same position that causes a  transition.
transition.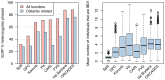
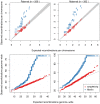
 ) are removed (blue). Right: The distributions of the average number of “surrogate” parents for each cohort when closely related pairs (
) are removed (blue). Right: The distributions of the average number of “surrogate” parents for each cohort when closely related pairs ( ) are removed.
) are removed.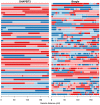
 or
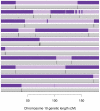
or  transition, both of which imply a SE in the child. Changes of colour between light and dark blue or between light and dark red correspond to
transition, both of which imply a SE in the child. Changes of colour between light and dark blue or between light and dark red correspond to  transitions, which correspond to a change on IBD state in the parent, and could be caused by a recombination or a SE in the parent. The x-axis shows the sex-averaged genetic distance across the chromosome in centiMorgans.
transitions, which correspond to a change on IBD state in the parent, and could be caused by a recombination or a SE in the parent. The x-axis shows the sex-averaged genetic distance across the chromosome in centiMorgans.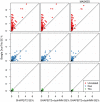
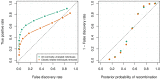
 ). Left: The ROC curves for recombination detection in uninformative duos for our duo HMM using the SHAPEIT2 haplotypes. Right: The average number of correct detections against the average posterior probability. Setting a high probability threshold ensures a very low false discovery rate.
). Left: The ROC curves for recombination detection in uninformative duos for our duo HMM using the SHAPEIT2 haplotypes. Right: The average number of correct detections against the average posterior probability. Setting a high probability threshold ensures a very low false discovery rate.References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

