Reactivation of maternal SNORD116 cluster via SETDB1 knockdown in Prader-Willi syndrome iPSCs
- PMID: 24760766
- PMCID: PMC4481691
- DOI: 10.1093/hmg/ddu187
Reactivation of maternal SNORD116 cluster via SETDB1 knockdown in Prader-Willi syndrome iPSCs
Abstract
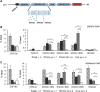
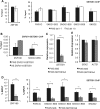
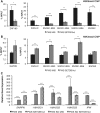
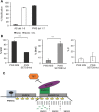
Prader-Willi syndrome (PWS), a disorder of genomic imprinting, is characterized by neonatal hypotonia, hypogonadism, small hands and feet, hyperphagia and obesity in adulthood. PWS results from the loss of paternal copies of the cluster of SNORD116 C/D box snoRNAs and their host transcript, 116HG, on human chromosome 15q11-q13. We have investigated the mechanism of repression of the maternal SNORD116 cluster and 116HG. Here, we report that the zinc-finger protein ZNF274, in association with the histone H3 lysine 9 (H3K9) methyltransferase SETDB1, is part of a complex that binds to the silent maternal but not the active paternal alleles. Knockdown of SETDB1 in PWS-specific induced pluripotent cells (iPSCs) causes a decrease in the accumulation of H3K9 trimethylation (H3K9me3) at 116HG and corresponding accumulation of the active chromatin mark histone H3 lysine 4 dimethylation (H3K4me2). We also show that upon knockdown of SETDB1 in PWS-specific iPSCs, expression of maternally silenced 116HG RNA is partially restored. SETDB1 knockdown in PWS iPSCs also disrupts DNA methylation at the PWS-IC where a decrease in 5-methylcytosine is observed in association with a concomitant increase in 5-hydroxymethylcytosine. This observation suggests that the ZNF274/SETDB1 complex bound to the SNORD116 cluster may protect the PWS-IC from DNA demethylation during early development. Our findings reveal novel epigenetic mechanisms that function to repress the maternal 15q11-q13 region.
© The Author 2014. Published by Oxford University Press. All rights reserved. For Permissions, please email: journals.permissions@oup.com.
Figures

Similar articles
-
Specific ZNF274 binding interference at SNORD116 activates the maternal transcripts in Prader-Willi syndrome neurons.Hum Mol Genet. 2020 Nov 25;29(19):3285-3295. doi: 10.1093/hmg/ddaa210. Hum Mol Genet. 2020. PMID: 32977341 Free PMC article.
-
Zinc finger protein 274 regulates imprinted expression of transcripts in Prader-Willi syndrome neurons.Hum Mol Genet. 2018 Feb 1;27(3):505-515. doi: 10.1093/hmg/ddx420. Hum Mol Genet. 2018. PMID: 29228278
-
RNAi Knockdown of EHMT2 in Maternal Expression of Prader-Willi Syndrome Genes.Genes (Basel). 2024 Oct 24;15(11):1366. doi: 10.3390/genes15111366. Genes (Basel). 2024. PMID: 39596566 Free PMC article.
-
Epigenetic therapy of Prader-Willi syndrome.Transl Res. 2019 Jun;208:105-118. doi: 10.1016/j.trsl.2019.02.012. Epub 2019 Mar 5. Transl Res. 2019. PMID: 30904443 Free PMC article. Review.
-
Cognitive deficits in the Snord116 deletion mouse model for Prader-Willi syndrome.Neurobiol Learn Mem. 2019 Nov;165:106874. doi: 10.1016/j.nlm.2018.05.011. Epub 2018 May 23. Neurobiol Learn Mem. 2019. PMID: 29800646 Free PMC article. Review.
Cited by
-
CRISPR/Cas9 Epigenome Editing Potential for Rare Imprinting Diseases: A Review.Cells. 2020 Apr 16;9(4):993. doi: 10.3390/cells9040993. Cells. 2020. PMID: 32316223 Free PMC article. Review.
-
Promising therapeutic aspects in human genetic imprinting disorders.Clin Epigenetics. 2022 Nov 12;14(1):146. doi: 10.1186/s13148-022-01369-6. Clin Epigenetics. 2022. PMID: 36371218 Free PMC article. Review.
-
Specific ZNF274 binding interference at SNORD116 activates the maternal transcripts in Prader-Willi syndrome neurons.Hum Mol Genet. 2020 Nov 25;29(19):3285-3295. doi: 10.1093/hmg/ddaa210. Hum Mol Genet. 2020. PMID: 32977341 Free PMC article.
-
T Cell Transcriptomes Describe Patient Subtypes in Systemic Lupus Erythematosus.PLoS One. 2015 Nov 6;10(11):e0141171. doi: 10.1371/journal.pone.0141171. eCollection 2015. PLoS One. 2015. PMID: 26544975 Free PMC article.
-
IPSC Models of Chromosome 15Q Imprinting Disorders: From Disease Modeling to Therapeutic Strategies.Adv Neurobiol. 2020;25:55-77. doi: 10.1007/978-3-030-45493-7_3. Adv Neurobiol. 2020. PMID: 32578144 Free PMC article.
References
-
- Cassidy S.B., Schwartz S., Miller J.L., Driscoll D.J. Prader-Willi syndrome. Genet. Med. 2012;14:10–26. doi:10.1038/gim.1030b1013e31822bead31820. - DOI - PubMed
-
- Saitoh S., Buiting K., Rogan P.K., Buxton J.L., Driscoll D.J., Arnemann J., Konig R., Malcolm S., Horsthemke B., Nicholls R.D. Minimal definition of the imprinting center and fixation of chromosome 15q11-q13 epigenotype by imprinting mutations. Proc. Natl. Acad. Sci. USA. 1996;93:7811–7815. - PMC - PubMed
-
- Boccaccio I., Glatt-Deeley H., Watrin F., Roeckel N., Lalande M., Muscatelli F. The human MAGEL2 gene and its mouse homologue are paternally expressed and mapped to the Prader-Willi region. Hum. Mol. Genet. 1999;8:2497–2505. - PubMed
-
- Jay P., Rougeulle C., Massacrier A., Moncla A., Mattei M.G., Malzac P., Roeckel N., Taviaux S., Lefranc J.L., Cau P., et al. The human necdin gene, NDN, is maternally imprinted and located in the Prader-Willi syndrome chromosomal region. Nat. Genet. 1997;17:357–361. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

