Targeted CNS Delivery Using Human MiniPromoters and Demonstrated Compatibility with Adeno-Associated Viral Vectors
- PMID: 24761428
- PMCID: PMC3992516
- DOI: 10.1038/mtm.2013.5
Targeted CNS Delivery Using Human MiniPromoters and Demonstrated Compatibility with Adeno-Associated Viral Vectors
Abstract
Critical for human gene therapy is the availability of small promoter tools to drive gene expression in a highly specific and reproducible manner. We tackled this challenge by developing human DNA MiniPromoters using computational biology and phylogenetic conservation. MiniPromoters were tested in mouse as single-copy knock-ins at the Hprt locus on the X Chromosome, and evaluated for lacZ reporter expression in CNS and non-CNS tissue. Eighteen novel MiniPromoters driving expression in mouse brain were identified, two MiniPromoters for driving pan-neuronal expression, and 17 MiniPromoters for the mouse eye. Key areas of therapeutic interest were represented in this set: the cerebral cortex, embryonic hypothalamus, spinal cord, bipolar and ganglion cells of the retina, and skeletal muscle. We also demonstrated that three retinal ganglion cell MiniPromoters exhibit similar cell-type specificity when delivered via adeno-associated virus (AAV) vectors intravitreally. We conclude that our methodology and characterization has resulted in desirable expression characteristics that are intrinsic to the MiniPromoter, not dictated by copy number effects or genomic location, and results in constructs predisposed to success in AAV. These MiniPromoters are immediately applicable for pre-clinical studies towards gene therapy in humans, and are publicly available to facilitate basic and clinical research, and human gene therapy.
Keywords: AAV; gene therapy; promoters.
Figures
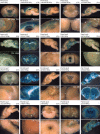
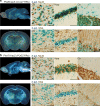

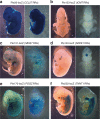



References
-
- Ng L, Bernard A, Lau C, Overly CC, Dong H-W, Kuan C. An anatomic gene expression atlas of the adult mouse brain. Nat Neurosci. 2009;12:356–362. - PubMed
-
- Gong S, Zheng C, Doughty ML, Losos K, Didkovsky N, Schambra UB. A gene expression atlas of the central nervous system based on bacterial artificial chromosomes. Nature. 2003;425:917–925. - PubMed
-
- Visel A, Thaller C, Eichele G. GenePaint.org: an atlas of gene expression patterns in the mouse embryo. Nucleic Acids Res. 2004;32 Database issue:D552–D556. - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

