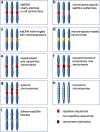
Centromere identity from the DNA point of view
- PMID: 24763964
- PMCID: PMC4107277
- DOI: 10.1007/s00412-014-0462-0
Centromere identity from the DNA point of view
Abstract
The centromere is a chromosomal locus responsible for the faithful segregation of genetic material during cell division. It has become evident that centromeres can be established literally on any DNA sequence, and the possible synergy between DNA sequences and the most prominent centromere identifiers, protein components, and epigenetic marks remains uncertain. However, some evolutionary preferences seem to exist, and long-term established centromeres are frequently formed on long arrays of satellite DNAs and/or transposable elements. Recent progress in understanding functional centromere sequences is based largely on the high-resolution DNA mapping of sequences that interact with the centromere-specific histone H3 variant, the most reliable marker of active centromeres. In addition, sequence assembly and mapping of large repetitive centromeric regions, as well as comparative genome analyses offer insight into their complex organization and evolution. The rapidly advancing field of transcription in centromere regions highlights the functional importance of centromeric transcripts. Here, we comprehensively review the current state of knowledge on the composition and functionality of DNA sequences underlying active centromeres and discuss their contribution to the functioning of different centromere types in higher eukaryotes.
Figures
References
-
- Bachmann L, Raab M, Sperlich D. Satellite DNA and speciation—a species-specific satellite DNA of Drosophila guanche. J Zool Syst Evol Res. 1989;27:84–93.
-
- Black BE, Bassett EA. The histone variant CENP-A and centromere specification. Curr Opin Cell Biol. 2008;20:91–100. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources