iMODS: internal coordinates normal mode analysis server
- PMID: 24771341
- PMCID: PMC4086069
- DOI: 10.1093/nar/gku339
iMODS: internal coordinates normal mode analysis server
Abstract
Normal mode analysis (NMA) in internal (dihedral) coordinates naturally reproduces the collective functional motions of biological macromolecules. iMODS facilitates the exploration of such modes and generates feasible transition pathways between two homologous structures, even with large macromolecules. The distinctive internal coordinate formulation improves the efficiency of NMA and extends its applicability while implicitly maintaining stereochemistry. Vibrational analysis, motion animations and morphing trajectories can be easily carried out at different resolution scales almost interactively. The server is versatile; non-specialists can rapidly characterize potential conformational changes, whereas advanced users can customize the model resolution with multiple coarse-grained atomic representations and elastic network potentials. iMODS supports advanced visualization capabilities for illustrating collective motions, including an improved affine-model-based arrow representation of domain dynamics. The generated all-heavy-atoms conformations can be used to introduce flexibility for more advanced modeling or sampling strategies. The server is free and open to all users with no login requirement at http://imods.chaconlab.org.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures
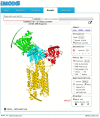
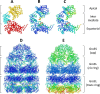

References
-
- Dykeman E.C., Sankey O.F. Normal mode analysis and applications in biological physics. J. Phys. Condens. Matter. 2010;22:423202. - PubMed
-
- Ma J. Usefulness and limitations of normal mode analysis in modeling dynamics of biomolecular complexes. Structure. 2005;13:373–380. - PubMed
-
- Skjaerven L., Hollup S.M., Reuter N. Normal mode analysis for proteins. J. Mol. Struct.: THEOCHEM. 2009;898:42–48.
-
- Zheng W., Brooks B.R., Hummer G. Protein conformational transitions explored by mixed elastic network models. Proteins: Struct. Funct. Bioinformatics. 2007;69:43–57. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

