Somatic mutation as a mechanism of Wnt/β-catenin pathway activation in CLL
- PMID: 24778153
- PMCID: PMC4133483
- DOI: 10.1182/blood-2014-01-552067
Somatic mutation as a mechanism of Wnt/β-catenin pathway activation in CLL
Abstract
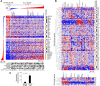
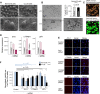
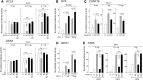
One major goal of cancer genome sequencing is to identify key genes and pathways that drive tumor pathogenesis. Although many studies have identified candidate driver genes based on recurrence of mutations in individual genes, subsets of genes with nonrecurrent mutations may also be defined as putative drivers if they affect a single biological pathway. In this fashion, we previously identified Wnt signaling as significantly mutated through large-scale massively parallel DNA sequencing of chronic lymphocytic leukemia (CLL). Here, we use a novel method of biomolecule delivery, vertical silicon nanowires, to efficiently introduce small interfering RNAs into CLL cells, and interrogate the effects of 8 of 15 mutated Wnt pathway members identified across 91 CLLs. In HEK293T cells, mutations in 2 genes did not generate functional changes, 3 led to dysregulated pathway activation, and 3 led to further activation or loss of repression of pathway activation. Silencing 4 of 8 mutated genes in CLL samples harboring the mutated alleles resulted in reduced viability compared with leukemia samples with wild-type alleles. We demonstrate that somatic mutations in CLL can generate dependence on this pathway for survival. These findings support the notion that nonrecurrent mutations at different nodes of the Wnt pathway can contribute to leukemogenesis.
© 2014 by The American Society of Hematology.
Figures



Comment in
-
Old and new news in CLL: "It's the pathway, stupid!".Blood. 2014 Aug 14;124(7):989-90. doi: 10.1182/blood-2014-05-574145. Blood. 2014. PMID: 25124780 No abstract available.
References
-
- Quesada V, Conde L, Villamor N, et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat Genet. 2012;44(1):47–52. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

