Allele-specific genome-wide profiling in human primary erythroblasts reveal replication program organization
- PMID: 24787348
- PMCID: PMC4006724
- DOI: 10.1371/journal.pgen.1004319
Allele-specific genome-wide profiling in human primary erythroblasts reveal replication program organization
Abstract
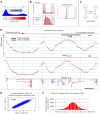
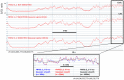
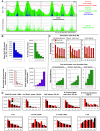
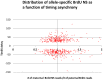
We have developed a new approach to characterize allele-specific timing of DNA replication genome-wide in human primary basophilic erythroblasts. We show that the two chromosome homologs replicate at the same time in about 88% of the genome and that large structural variants are preferentially associated with asynchronous replication. We identified about 600 megabase-sized asynchronously replicated domains in two tested individuals. The longest asynchronously replicated domains are enriched in imprinted genes suggesting that structural variants and parental imprinting are two causes of replication asynchrony in the human genome. Biased chromosome X inactivation in one of the two individuals tested was another source of detectable replication asynchrony. Analysis of high-resolution TimEX profiles revealed small variations termed timing ripples, which were undetected in previous, lower resolution analyses. Timing ripples reflect highly reproducible, variations of the timing of replication in the 100 kb-range that exist within the well-characterized megabase-sized replication timing domains. These ripples correspond to clusters of origins of replication that we detected using novel nascent strands DNA profiling methods. Analysis of the distribution of replication origins revealed dramatic differences in initiation of replication frequencies during S phase and a strong association, in both synchronous and asynchronous regions, between origins of replication and three genomic features: G-quadruplexes, CpG Islands and transcription start sites. The frequency of initiation in asynchronous regions was similar in the two homologs. Asynchronous regions were richer in origins of replication than synchronous regions.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Aladjem MI (2007) Replication in context: dynamic regulation of DNA replication patterns in metazoans. Nat Rev Genet 8: 588–600. - PubMed
-
- Masai H, Matsumoto S, You Z, Yoshizawa-Sugata N, Oda M (2010) Eukaryotic chromosome DNA replication: where, when, and how? Annu Rev Biochem 79: 89–130. - PubMed
-
- Mechali M (2010) Eukaryotic DNA replication origins: many choices for appropriate answers. Nat Rev Mol Cell Biol 11: 728–738. - PubMed
-
- Mechali M (2001) DNA replication origins: from sequence specificity to epigenetics. Nat Rev Genet 2: 640–645. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

