Linking activity and function to ecosystem dynamics in a coastal bacterioplankton community
- PMID: 24795712
- PMCID: PMC4006046
- DOI: 10.3389/fmicb.2014.00185
Linking activity and function to ecosystem dynamics in a coastal bacterioplankton community
Abstract
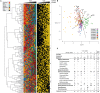
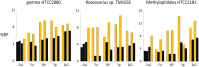
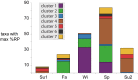
For bacterial communities containing hundreds to thousands of distinct populations, connecting functional processes and environmental dynamics at high taxonomic resolution has remained challenging. Here we use the expression of ribosomal proteins (%RP) as a proxy for in situ activity of 200 taxa within 20 metatranscriptomic samples in a coastal ocean time series encompassing both seasonal variability and diel dynamics. %RP patterns grouped the taxa into seven activity clusters with distinct profiles in functional gene expression and correlations with environmental gradients. Clusters 1-3 had their highest potential activity in the winter and fall, and included some of the most active taxa, while Clusters 4-7 had their highest potential activity in the spring and summer. Cluster 1 taxa were characterized by gene expression for motility and complex carbohydrate degradation (dominated by Gammaproteobacteria and Bacteroidetes), and Cluster 2 taxa by transcription of genes for amino acid and aromatic compound metabolism and aerobic anoxygenic phototrophy (Roseobacter). Other activity clusters were enriched in transcripts for proteorhodopsin and methylotrophy (Cluster 4; SAR11 and methylotrophs), photosynthesis and attachment (Clusters 5 and 7; Synechococcus, picoeukaryotes, Verucomicrobia, and Planctomycetes), and sulfur oxidation (Cluster 7; Gammaproteobacteria). The seasonal patterns in activity were overlain, and sometimes obscured, by large differences in %RP over shorter day-night timescales. Seventy-eight taxa, many of them heterotrophs, had a higher %RP activity index during the day than night, indicating a strong diel activity rhythm at this coastal site. Emerging from these taxonomically- and time-resolved estimates of in situ microbial activity are predictions of specific ecological groupings of microbial taxa in a dynamic coastal environment.
Keywords: activity; bacterioplankton; diel; marine; metatranscriptomics; seasonal.
Figures



Similar articles
-
A High-Resolution Time Series Reveals Distinct Seasonal Patterns of Planktonic Fungi at a Temperate Coastal Ocean Site (Beaufort, North Carolina, USA).Appl Environ Microbiol. 2018 Oct 17;84(21):e00967-18. doi: 10.1128/AEM.00967-18. Print 2018 Nov 1. Appl Environ Microbiol. 2018. PMID: 30143506 Free PMC article.
-
Metatranscriptomic signature of exogenous polyamine utilization by coastal bacterioplankton.Environ Microbiol Rep. 2011 Dec;3(6):798-806. doi: 10.1111/j.1758-2229.2011.00289.x. Epub 2011 Sep 19. Environ Microbiol Rep. 2011. PMID: 23761372
-
Seasonal and spatial dynamics of bacterioplankton communities in a brackish water coastal lagoon.Sci Total Environ. 2020 Feb 25;705:134729. doi: 10.1016/j.scitotenv.2019.134729. Epub 2019 Dec 5. Sci Total Environ. 2020. PMID: 31838414
-
Strong Seasonality in Arctic Estuarine Microbial Food Webs.Front Microbiol. 2019 Nov 29;10:2628. doi: 10.3389/fmicb.2019.02628. eCollection 2019. Front Microbiol. 2019. PMID: 31849850 Free PMC article. Review.
-
The Minderoo-Monaco Commission on Plastics and Human Health.Ann Glob Health. 2023 Mar 21;89(1):23. doi: 10.5334/aogh.4056. eCollection 2023. Ann Glob Health. 2023. PMID: 36969097 Free PMC article. Review.
Cited by
-
Microbial Surface Colonization and Biofilm Development in Marine Environments.Microbiol Mol Biol Rev. 2015 Dec 23;80(1):91-138. doi: 10.1128/MMBR.00037-15. Print 2016 Mar. Microbiol Mol Biol Rev. 2015. PMID: 26700108 Free PMC article. Review.
-
Fine grained compositional analysis of Port Everglades Inlet microbiome using high throughput DNA sequencing.PeerJ. 2018 May 8;6:e4671. doi: 10.7717/peerj.4671. eCollection 2018. PeerJ. 2018. PMID: 29761039 Free PMC article.
-
Transcriptional Patterns of Biogeochemically Relevant Marker Genes by Temperate Marine Bacteria.Front Microbiol. 2020 Mar 20;11:465. doi: 10.3389/fmicb.2020.00465. eCollection 2020. Front Microbiol. 2020. PMID: 32265888 Free PMC article.
-
Transcriptional activity differentiates families of Marine Group II Euryarchaeota in the coastal ocean.ISME Commun. 2021 Mar 22;1(1):5. doi: 10.1038/s43705-021-00002-6. ISME Commun. 2021. PMID: 37938231 Free PMC article.
-
Causes and Consequences of a Variant Strain of Phaeobacter inhibens With Reduced Competition.Front Microbiol. 2018 Nov 2;9:2601. doi: 10.3389/fmicb.2018.02601. eCollection 2018. Front Microbiol. 2018. PMID: 30450086 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Other Literature Sources

