Epigenetic Regulation of Individual Modules of the immunoglobulin heavy chain locus 3' Regulatory Region
- PMID: 24795714
- PMCID: PMC4000994
- DOI: 10.3389/fimmu.2014.00163
Epigenetic Regulation of Individual Modules of the immunoglobulin heavy chain locus 3' Regulatory Region
Abstract
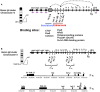
The Igh locus undergoes an amazing array of DNA rearrangements and modifications during B cell development. During early stages, the variable region gene is constructed from constituent variable (V), diversity (D), and joining (J) segments (VDJ joining). B cells that successfully express an antibody can be activated, leading to somatic hypermutation (SHM) focused on the variable region, and class switch recombination (CSR), which substitutes downstream constant region genes for the originally used Cμ constant region gene. Many investigators, ourselves included, have sought to understand how these processes specifically target the Igh locus and avoid other loci and potential deleterious consequences of malignant transformation. Our laboratory has concentrated on a complex regulatory region (RR) that is located downstream of Cα, the most 3' of the Igh constant region genes. The ~40 kb 3' RR, which is predicted to serve as a downstream major regulator of the Igh locus, contains two distinct segments: an ~28 kb region comprising four enhancers, and an adjacent ~12 kb region containing multiple CTCF and Pax5 binding sites. Analysis of targeted mutations in mice by a number of investigators has concluded that the entire 3' RR enhancer region is essential for SHM and CSR (but not for VDJ joining) and for high levels of expression of multiple isotypes. The CTCF/Pax5 binding region is a candidate for influencing VDJ joining early in B cell development and serving as a potential insulator of the Igh locus. Components of the 3' RR are subject to a variety of epigenetic changes during B cell development, i.e., DNAse I hypersensitivity, histone modifications, and DNA methylation, in association with transcription factor binding. I propose that these changes provide a foundation by which regulatory elements in modules of the 3' RR function by interacting with each other and with target sequences of the Igh locus.
Keywords: CTCF; Pax5; class switch recombination; enhancers; immunoglobulin heavy chain gene locus; insulators; somatic hypermutation.
Figures
Similar articles
-
The role of CTCF binding sites in the 3' immunoglobulin heavy chain regulatory region.Front Genet. 2012 Nov 16;3:251. doi: 10.3389/fgene.2012.00251. eCollection 2012. Front Genet. 2012. PMID: 23162572 Free PMC article.
-
Elucidation of a downstream boundary of the 3' IgH regulatory region.Mol Immunol. 2003 Jan;39(12):753-60. doi: 10.1016/s0161-5890(02)00256-0. Mol Immunol. 2003. PMID: 12531286
-
Interaction between the immunoglobulin heavy chain 3' regulatory region and the IgH transcription unit during B cell differentiation.Mol Immunol. 2011 Oct;49(1-2):297-303. doi: 10.1016/j.molimm.2011.08.024. Epub 2011 Sep 25. Mol Immunol. 2011. PMID: 21945019 Free PMC article.
-
Mechanism and control of V(D)J recombination versus class switch recombination: similarities and differences.Adv Immunol. 2005;86:43-112. doi: 10.1016/S0065-2776(04)86002-4. Adv Immunol. 2005. PMID: 15705419 Review.
-
The IgH locus 3' regulatory region: pulling the strings from behind.Adv Immunol. 2011;110:27-70. doi: 10.1016/B978-0-12-387663-8.00002-8. Adv Immunol. 2011. PMID: 21762815 Review.
Cited by
-
Chromatin and transcriptional tango on the immune dance floor.Front Immunol. 2014 Dec 15;5:631. doi: 10.3389/fimmu.2014.00631. eCollection 2014. Front Immunol. 2014. PMID: 25566246 Free PMC article. No abstract available.
-
Noncoding RNA processing by DIS3 regulates chromosomal architecture and somatic hypermutation in B cells.Nat Genet. 2021 Feb;53(2):230-242. doi: 10.1038/s41588-020-00772-0. Epub 2021 Feb 1. Nat Genet. 2021. PMID: 33526923 Free PMC article.
-
RNA exosome-regulated long non-coding RNA transcription controls super-enhancer activity.Cell. 2015 May 7;161(4):774-89. doi: 10.1016/j.cell.2015.04.034. Cell. 2015. PMID: 25957685 Free PMC article.
-
The 3'-Jα Region of the TCRα Locus Bears Gene Regulatory Activity in Thymic and Peripheral T Cells.PLoS One. 2015 Jul 15;10(7):e0132856. doi: 10.1371/journal.pone.0132856. eCollection 2015. PLoS One. 2015. PMID: 26177549 Free PMC article.
-
Nuclear Receptors, Ligands and the Mammalian B Cell.Int J Mol Sci. 2020 Jul 15;21(14):4997. doi: 10.3390/ijms21144997. Int J Mol Sci. 2020. PMID: 32679815 Free PMC article. Review.
References
-
- Vincent-Fabert C, Fiancette R, Pinaud E, Truffinet V, Cogne N, Cogne M, et al. Genomic deletion of the whole Igh 3’ regulatory region (hs3a, hs1,2, hs3b, hs4) dramatically affects class switch recombination and Ig secretion to all isotypes. Blood (2010) 116:1895–810.1182/blood-2010-01-264689 - DOI - PubMed
-
- Michaelson JS, Singh M, Snapper CM, Sha WC, Baltimore D, Birshtein BK. Regulation of 3’ Igh enhancers by a common set of factors, including kappa B-binding proteins. J Immunol (1996) 156:2828–39 - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

