Asymmetric localization of natural antisense RNA of neuropeptide sensorin in Aplysia sensory neurons during aging and activity
- PMID: 24795747
- PMCID: PMC4001032
- DOI: 10.3389/fgene.2014.00084
Asymmetric localization of natural antisense RNA of neuropeptide sensorin in Aplysia sensory neurons during aging and activity
Abstract
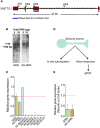
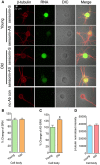
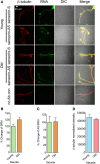
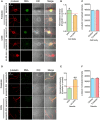
Despite the advances in our understanding of transcriptome, regulation and function of its non-coding components continue to be poorly understood. Here we searched for natural antisense transcript for sensorin (NAT-SRN), a neuropeptide expressed in the presynaptic sensory neurons of gill-withdrawal reflex of the marine snail Aplysia californica. Sensorin (SRN) has a key role in learning and long-term memory storage in Aplysia. We have now identified NAT-SRN in the central nervous system (CNS) and have confirmed its expression by northern blotting and fluorescent RNA in situ hybridization. Quantitative analysis of NAT-SRN in micro-dissected cell bodies and processes of sensory neurons suggest that NAT-SRN is present in the distal neuronal processes along with sense transcripts. Importantly, aging is associated with reduction in levels of NAT-SRN in sensory neuron processes. Furthermore, we find that forskolin, an activator of CREB signaling, differentially alters the distribution of SRN and NAT-SRN. These studies reveal novel insights into physiological regulation of natural antisense RNAs.
Keywords: Aplysia; aging; antisense RNA; memory; neural circuitry; nocoding RNA.
Figures
Similar articles
-
Comparative localization of two serotonin receptors and sensorin in the central nervous system of Aplysia californica.J Comp Neurol. 2005 Sep 26;490(3):295-304. doi: 10.1002/cne.20666. J Comp Neurol. 2005. PMID: 16082675
-
Whole-transcriptome changes in gene expression accompany aging of sensory neurons in Aplysia californica.BMC Genomics. 2018 Jul 11;19(1):529. doi: 10.1186/s12864-018-4909-1. BMC Genomics. 2018. PMID: 29996779 Free PMC article.
-
Synapse formation and mRNA localization in cultured Aplysia neurons.Neuron. 2006 Feb 2;49(3):349-56. doi: 10.1016/j.neuron.2005.12.029. Neuron. 2006. PMID: 16446139
-
Transcriptional regulation of long-term memory in the marine snail Aplysia.Mol Brain. 2008 Jun 17;1:3. doi: 10.1186/1756-6606-1-3. Mol Brain. 2008. PMID: 18803855 Free PMC article. Review.
-
Postsynaptic regulation of the development and long-term plasticity of Aplysia sensorimotor synapses in cell culture.J Neurobiol. 1994 Jun;25(6):666-93. doi: 10.1002/neu.480250608. J Neurobiol. 1994. PMID: 8071666 Review.
Cited by
-
The seeds of its regulation: Natural antisense transcripts as single-gene control switches in neurodegenerative disorders.Ageing Res Rev. 2024 Aug;99:102336. doi: 10.1016/j.arr.2024.102336. Epub 2024 May 11. Ageing Res Rev. 2024. PMID: 38740308 Free PMC article. Review.
-
Transcriptome analyses of adult mouse brain reveal enrichment of lncRNAs in specific brain regions and neuronal populations.Front Cell Neurosci. 2015 Mar 6;9:63. doi: 10.3389/fncel.2015.00063. eCollection 2015. Front Cell Neurosci. 2015. PMID: 25798087 Free PMC article.
-
Out of darkness: long non-coding RNAs come of age.Front Genet. 2014 Nov 7;5:388. doi: 10.3389/fgene.2014.00388. eCollection 2014. Front Genet. 2014. PMID: 25426139 Free PMC article. No abstract available.
-
Long Non-Coding RNAs in Neuronal Aging.Noncoding RNA. 2018 Apr 18;4(2):12. doi: 10.3390/ncrna4020012. Noncoding RNA. 2018. PMID: 29670042 Free PMC article. Review.
-
Innovative perspectives on glioblastoma: the emerging role of long non-coding RNAs.Funct Integr Genomics. 2025 Feb 24;25(1):43. doi: 10.1007/s10142-025-01557-6. Funct Integr Genomics. 2025. PMID: 39992471 Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

