Wiring miRNAs to pathways: a topological approach to integrate miRNA and mRNA expression profiles
- PMID: 24803669
- PMCID: PMC4066781
- DOI: 10.1093/nar/gku354
Wiring miRNAs to pathways: a topological approach to integrate miRNA and mRNA expression profiles
Abstract
The production rate of gene expression data is nothing less than astounding. However, with the benefit of hindsight we can assert that, since we completely ignored the non-coding part of the transcriptome, we spent the last decade to study cell mechanisms having few data in our hands. In this scenario, microRNAs, which are key post-trascriptional regulators, deserve special attention. Given the state of knowledge about their biogenesis, mechanisms of action and the numerous experimentally validated target genes, miRNAs are also gradually appearing in the formal pathway representations such as KEGG and Reactome maps. However, the number of miRNAs annotated in pathway maps are very few and pathway analyses exploiting this new regulatory layer are still lacking. To fill these gaps, we present 'micrographite' a new pipeline to perform topological pathway analysis integrating gene and miRNA expression profiles. Here, micrographite is used to study and dissect the epithelial ovarian cancer gene and miRNA transcriptome defining and validating a new regulatory circuit related to ovarian cancer histotype specificity.
© The Author(s) 2014. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

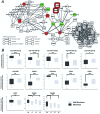
Similar articles
-
MicroRNA expression and gene regulation drive breast cancer progression and metastasis in PyMT mice.Breast Cancer Res. 2016 Jul 22;18(1):75. doi: 10.1186/s13058-016-0735-z. Breast Cancer Res. 2016. PMID: 27449149 Free PMC article.
-
Identification of miRNA-mRNA crosstalk in pancreatic cancer by integrating transcriptome analysis.Eur Rev Med Pharmacol Sci. 2015;19(5):825-34. Eur Rev Med Pharmacol Sci. 2015. PMID: 25807437
-
Construction and analysis of mRNA, miRNA, lncRNA, and TF regulatory networks reveal the key genes associated with prostate cancer.PLoS One. 2018 Aug 23;13(8):e0198055. doi: 10.1371/journal.pone.0198055. eCollection 2018. PLoS One. 2018. PMID: 30138363 Free PMC article.
-
Integrated analysis of microRNA and gene expression profiles reveals a functional regulatory module associated with liver fibrosis.Gene. 2017 Dec 15;636:87-95. doi: 10.1016/j.gene.2017.09.027. Epub 2017 Sep 14. Gene. 2017. PMID: 28919164
-
Identifying miRNA-mediated signaling subpathways by integrating paired miRNA/mRNA expression data with pathway topology.Annu Int Conf IEEE Eng Med Biol Soc. 2015;2015:3997-4000. doi: 10.1109/EMBC.2015.7319270. Annu Int Conf IEEE Eng Med Biol Soc. 2015. PMID: 26737170
Cited by
-
Computational Oncology in the Multi-Omics Era: State of the Art.Front Oncol. 2020 Apr 7;10:423. doi: 10.3389/fonc.2020.00423. eCollection 2020. Front Oncol. 2020. PMID: 32318338 Free PMC article. Review.
-
A systems biology approach to investigate the mechanism of action of trabectedin in a model of myelomonocytic leukemia.Pharmacogenomics J. 2018 Jan;18(1):56-63. doi: 10.1038/tpj.2016.76. Epub 2016 Dec 13. Pharmacogenomics J. 2018. PMID: 27958379 Free PMC article.
-
A mitochondrial NADPH-cholesterol axis regulates extracellular vesicle biogenesis to support hematopoietic stem cell fate.Cell Stem Cell. 2024 Mar 7;31(3):359-377.e10. doi: 10.1016/j.stem.2024.02.004. Cell Stem Cell. 2024. PMID: 38458178 Free PMC article.
-
Disentangling the microRNA regulatory milieu in multiple myeloma: integrative genomics analysis outlines mixed miRNA-TF circuits and pathway-derived networks modulated in t(4;14) patients.Oncotarget. 2016 Jan 19;7(3):2367-78. doi: 10.18632/oncotarget.6151. Oncotarget. 2016. PMID: 26496024 Free PMC article.
-
Circulating Cell-Free Nucleic Acids as Epigenetic Biomarkers in Precision Medicine.Front Genet. 2020 Aug 11;11:844. doi: 10.3389/fgene.2020.00844. eCollection 2020. Front Genet. 2020. PMID: 32849827 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

