Mechanical operation and intersubunit coordination of ring-shaped molecular motors: insights from single-molecule studies
- PMID: 24806916
- PMCID: PMC4017299
- DOI: 10.1016/j.bpj.2014.03.029
Mechanical operation and intersubunit coordination of ring-shaped molecular motors: insights from single-molecule studies
Abstract
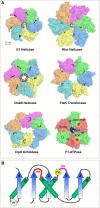
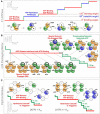
Ring NTPases represent a large and diverse group of proteins that couple their nucleotide hydrolysis activity to a mechanical task involving force generation and some type of transport process in the cell. Because of their shape, these enzymes often operate as gates that separate distinct cellular compartments to control and regulate the passage of chemical species across them. In this manner, ions and small molecules are moved across membranes, biopolymer substrates are segregated between cells or moved into confined spaces, double-stranded nucleic acids are separated into single strands to provide access to the genetic information, and polypeptides are unfolded and processed for recycling. Here we review the recent advances in the characterization of these motors using single-molecule manipulation and detection approaches. We describe the various mechanisms by which ring motors convert chemical energy to mechanical force or torque and coordinate the activities of individual subunits that constitute the ring. We also examine how single-molecule studies have contributed to a better understanding of the structural elements involved in motor-substrate interaction, mechanochemical coupling, and intersubunit coordination. Finally, we discuss how these molecular motors tailor their operation-often through regulation by other cofactors-to suit their unique biological functions.
Copyright © 2014 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures


Similar articles
-
Intersubunit coordination and cooperativity in ring-shaped NTPases.Curr Opin Struct Biol. 2013 Apr;23(2):229-34. doi: 10.1016/j.sbi.2013.01.004. Epub 2013 Feb 8. Curr Opin Struct Biol. 2013. PMID: 23395511 Review.
-
Coordination and control inside simple biomolecular machines.Adv Exp Med Biol. 2014;805:353-84. doi: 10.1007/978-3-319-02970-2_15. Adv Exp Med Biol. 2014. PMID: 24446369 Review.
-
Mechanochemical coupling and bi-phasic force-velocity dependence in the ultra-fast ring ATPase SpoIIIE.Elife. 2018 Mar 5;7:e32354. doi: 10.7554/eLife.32354. Elife. 2018. PMID: 29504934 Free PMC article.
-
Subunits interactions in kinesin motors.Eur J Cell Biol. 2007 Sep;86(9):559-68. doi: 10.1016/j.ejcb.2007.05.008. Epub 2007 Jul 12. Eur J Cell Biol. 2007. PMID: 17628208 Review.
-
The nuts and bolts of ring-translocase structure and mechanism.Curr Opin Struct Biol. 2011 Apr;21(2):240-8. doi: 10.1016/j.sbi.2011.01.002. Epub 2011 Feb 1. Curr Opin Struct Biol. 2011. PMID: 21282052 Free PMC article. Review.
Cited by
-
NMR structure of a vestigial nuclease provides insight into the evolution of functional transitions in viral dsDNA packaging motors.Nucleic Acids Res. 2020 Nov 18;48(20):11737-11749. doi: 10.1093/nar/gkaa874. Nucleic Acids Res. 2020. PMID: 33089330 Free PMC article.
-
Molecular switch-like regulation in motor proteins.Philos Trans R Soc Lond B Biol Sci. 2018 Jun 19;373(1749):20170181. doi: 10.1098/rstb.2017.0181. Philos Trans R Soc Lond B Biol Sci. 2018. PMID: 29735735 Free PMC article. Review.
-
NMR Structures and Dynamics in a Prohead RNA Loop that Binds Metal Ions.J Phys Chem Lett. 2016 Oct 6;7(19):3841-3846. doi: 10.1021/acs.jpclett.6b01465. Epub 2016 Sep 19. J Phys Chem Lett. 2016. PMID: 27631837 Free PMC article.
-
Sequential allosteric mechanism of ATP hydrolysis by the CCT/TRiC chaperone is revealed through Arrhenius analysis.Proc Natl Acad Sci U S A. 2017 May 16;114(20):5189-5194. doi: 10.1073/pnas.1617746114. Epub 2017 May 1. Proc Natl Acad Sci U S A. 2017. PMID: 28461478 Free PMC article.
-
Assessing heterogeneity in oligomeric AAA+ machines.Cell Mol Life Sci. 2017 Mar;74(6):1001-1018. doi: 10.1007/s00018-016-2374-z. Epub 2016 Sep 26. Cell Mol Life Sci. 2017. PMID: 27669691 Free PMC article. Review.
References
-
- Bustamante C., Chemla Y.R., Izhaky D. Mechanical processes in biochemistry. Annu. Rev. Biochem. 2004;73:705–748. - PubMed
-
- Kolomeisky A.B., Fisher M.E. Molecular motors: a theorist’s perspective. Annu. Rev. Phys. Chem. 2007;58:675–695. - PubMed
-
- Iyer L.M., Leipe D.D., Aravind L. Evolutionary history and higher order classification of AAA+ ATPases. J. Struct. Biol. 2004;146:11–31. - PubMed
-
- Erzberger J.P., Berger J.M. Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu. Rev. Biophys. Biomol. Struct. 2006;35:93–114. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

