High-throughput platform for the discovery of elicitors of silent bacterial gene clusters
- PMID: 24808135
- PMCID: PMC4034229
- DOI: 10.1073/pnas.1400019111
High-throughput platform for the discovery of elicitors of silent bacterial gene clusters
Abstract
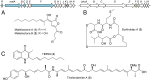
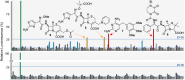
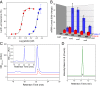
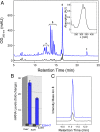
Over the past decade, bacterial genome sequences have revealed an immense reservoir of biosynthetic gene clusters, sets of contiguous genes that have the potential to produce drugs or drug-like molecules. However, the majority of these gene clusters appear to be inactive for unknown reasons prompting terms such as "cryptic" or "silent" to describe them. Because natural products have been a major source of therapeutic molecules, methods that rationally activate these silent clusters would have a profound impact on drug discovery. Herein, a new strategy is outlined for awakening silent gene clusters using small molecule elicitors. In this method, a genetic reporter construct affords a facile read-out for activation of the silent cluster of interest, while high-throughput screening of small molecule libraries provides potential inducers. This approach was applied to two cryptic gene clusters in the pathogenic model Burkholderia thailandensis. The results not only demonstrate a prominent activation of these two clusters, but also reveal that the majority of elicitors are themselves antibiotics, most in common clinical use. Antibiotics, which kill B. thailandensis at high concentrations, act as inducers of secondary metabolism at low concentrations. One of these antibiotics, trimethoprim, served as a global activator of secondary metabolism by inducing at least five biosynthetic pathways. Further application of this strategy promises to uncover the regulatory networks that activate silent gene clusters while at the same time providing access to the vast array of cryptic molecules found in bacteria.
Keywords: cryptic metabolites; malleilactone; natural products discovery.
Conflict of interest statement
The author declares no conflict of interest.
Figures
References
-
- Walsh C. Where will new antibiotics come from? Nat Rev Microbiol. 2003;1(1):65–70. - PubMed
-
- Nathan C. Antibiotics at the crossroads. Nature. 2004;431(7011):899–902. - PubMed
-
- Newman DJ, Cragg GM, Snader KM. The influence of natural products upon drug discovery. Nat Prod Rep. 2000;17(3):215–234. - PubMed
-
- Clatworthy AE, Pierson E, Hung DT. Targeting virulence: A new paradigm for antimicrobial therapy. Nat Chem Biol. 2007;3(9):541–548. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

