Phosphorylation of KasB regulates virulence and acid-fastness in Mycobacterium tuberculosis
- PMID: 24809459
- PMCID: PMC4014462
- DOI: 10.1371/journal.ppat.1004115
Phosphorylation of KasB regulates virulence and acid-fastness in Mycobacterium tuberculosis
Abstract
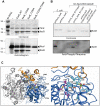
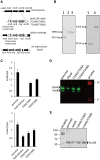
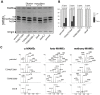
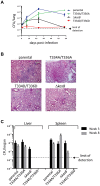
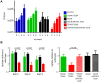
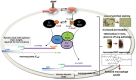
Mycobacterium tuberculosis bacilli display two signature features: acid-fast staining and the capacity to induce long-term latent infections in humans. However, the mechanisms governing these two important processes remain largely unknown. Ser/Thr phosphorylation has recently emerged as an important regulatory mechanism allowing mycobacteria to adapt their cell wall structure/composition in response to their environment. Herein, we evaluated whether phosphorylation of KasB, a crucial mycolic acid biosynthetic enzyme, could modulate acid-fast staining and virulence. Tandem mass spectrometry and site-directed mutagenesis revealed that phosphorylation of KasB occurred at Thr334 and Thr336 both in vitro and in mycobacteria. Isogenic strains of M. tuberculosis with either a deletion of the kasB gene or a kasB_T334D/T336D allele, mimicking constitutive phosphorylation of KasB, were constructed by specialized linkage transduction. Biochemical and structural analyses comparing these mutants to the parental strain revealed that both mutant strains had mycolic acids that were shortened by 4-6 carbon atoms and lacked trans-cyclopropanation. Together, these results suggested that in M. tuberculosis, phosphorylation profoundly decreases the condensing activity of KasB. Structural/modeling analyses reveal that Thr334 and Thr336 are located in the vicinity of the catalytic triad, which indicates that phosphorylation of these amino acids would result in loss of enzyme activity. Importantly, the kasB_T334D/T336D phosphomimetic and deletion alleles, in contrast to the kasB_T334A/T336A phosphoablative allele, completely lost acid-fast staining. Moreover, assessing the virulence of these strains indicated that the KasB phosphomimetic mutant was attenuated in both immunodeficient and immunocompetent mice following aerosol infection. This attenuation was characterized by the absence of lung pathology. Overall, these results highlight for the first time the role of Ser/Thr kinase-dependent KasB phosphorylation in regulating the later stages of mycolic acid elongation, with important consequences in terms of acid-fast staining and pathogenicity.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
-
- Zhou P, Long Q, Zhou Y, Wang H, Xie J (2012) Mycobacterium tuberculosis two-component systems and implications in novel vaccines and drugs. Crit Rev Eukaryot Gene Expr 22: 37–52. - PubMed
-
- Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, et al. (1998) Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature 393: 537–544. - PubMed
-
- Av-Gay Y, Everett M (2000) The eukaryotic-like Ser/Thr protein kinases of Mycobacterium tuberculosis. Trends Microbiol 8: 238–244. - PubMed
-
- Chao J, Wong D, Zheng X, Poirier V, Bach H, et al. (2010) Protein kinase and phosphatase signaling in Mycobacterium tuberculosis physiology and pathogenesis. Biochim Biophys Acta 1804: 620–627. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

