HupB, a nucleoid-associated protein of Mycobacterium tuberculosis, is modified by serine/threonine protein kinases in vivo
- PMID: 24816602
- PMCID: PMC4097597
- DOI: 10.1128/JB.01625-14
HupB, a nucleoid-associated protein of Mycobacterium tuberculosis, is modified by serine/threonine protein kinases in vivo
Abstract
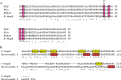
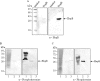
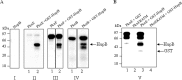
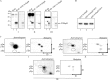
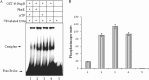
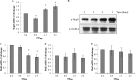
HU, a widely conserved bacterial histone-like protein, regulates many genes, including those involved in stress response and virulence. Whereas ample data are available on HU-DNA communication, the knowledge on how HU perceives a signal and transmit it to DNA remains limited. In this study, we identify HupB, the HU homolog of the human pathogen Mycobacterium tuberculosis, as a component of serine/threonine protein kinase (STPK) signaling. HupB is extracted in its native state from the exponentially growing cells of M. tuberculosis H37Ra and is shown to be phosphorylated on both serine and threonine residues. The STPKs capable of modifying HupB are determined in vitro and the residues modified by the STPKs are identified for both in vivo and the in vitro proteins through mass spectrometry. Of the identified phosphosites, Thr(65) and Thr(74) in the DNA-embracing β-strand of the N-terminal domain of HupB (N-HupB) are shown to be crucial for its interaction with DNA. In addition, Arg(55) is also identified as an important residue for N-HupB-DNA interaction. N-HupB is shown to have a diminished interaction with DNA after phosphorylation. Furthermore, hupB is shown to be maximally expressed during the stationary phase in M. tuberculosis H37Ra, while HupB kinases were found to be constitutively expressed (PknE and PknF) or most abundant during the exponential phase (PknB). In conclusion, HupB, a DNA-binding protein, with an ability to modulate chromatin structure is proposed to work in a growth-phase-dependent manner through its phosphorylation carried out by the mycobacterial STPKs.
Copyright © 2014, American Society for Microbiology. All Rights Reserved.
Figures


Similar articles
-
DNA clasping by mycobacterial HU: the C-terminal region of HupB mediates increased specificity of DNA binding.PLoS One. 2010 Sep 2;5(9):e12551. doi: 10.1371/journal.pone.0012551. PLoS One. 2010. PMID: 20824060 Free PMC article.
-
HupB Is a Bacterial Nucleoid-Associated Protein with an Indispensable Eukaryotic-Like Tail.mBio. 2017 Nov 7;8(6):e01272-17. doi: 10.1128/mBio.01272-17. mBio. 2017. PMID: 29114022 Free PMC article.
-
Synergy between the N-terminal and C-terminal domains of Mycobacterium tuberculosis HupB is essential for high-affinity binding, DNA supercoiling and inhibition of RecA-promoted strand exchange.FEBS J. 2011 Sep;278(18):3447-62. doi: 10.1111/j.1742-4658.2011.08267.x. Epub 2011 Aug 24. FEBS J. 2011. PMID: 21787377
-
Phosphoproteomic Approaches to Discover Novel Substrates of Mycobacterial Ser/Thr Protein Kinases.Mol Cell Proteomics. 2020 Feb;19(2):233-244. doi: 10.1074/mcp.R119.001668. Epub 2019 Dec 15. Mol Cell Proteomics. 2020. PMID: 31839597 Free PMC article. Review.
-
Mycobacterium tuberculosis Serine/Threonine Protein Kinases.Microbiol Spectr. 2014 Oct;2(5):10.1128/microbiolspec.MGM2-0006-2013. doi: 10.1128/microbiolspec.MGM2-0006-2013. Microbiol Spectr. 2014. PMID: 25429354 Free PMC article. Review.
Cited by
-
HupB, a nucleoid-associated protein, is critical for survival of Mycobacterium tuberculosis under host-mediated stresses and for enhanced tolerance to key first-line antibiotics.Front Microbiol. 2022 Aug 22;13:937970. doi: 10.3389/fmicb.2022.937970. eCollection 2022. Front Microbiol. 2022. PMID: 36071978 Free PMC article.
-
Molecular detection of M. tuberculosis and M. bovis and hematological and biochemical analyses in agricultural sprayers exposed to pesticides: A cross-sectional study in Punjab, Pakistan during 2014-2016.J Expo Sci Environ Epidemiol. 2017 Jul;27(4):434-443. doi: 10.1038/jes.2016.88. Epub 2017 Feb 1. J Expo Sci Environ Epidemiol. 2017. PMID: 28145400
-
Epigenetic Phosphorylation Control of Mycobacterium tuberculosis Infection and Persistence.Microbiol Spectr. 2017 Mar;5(2):10.1128/microbiolspec.tbtb2-0005-2015. doi: 10.1128/microbiolspec.TBTB2-0005-2015. Microbiol Spectr. 2017. PMID: 28281439 Free PMC article. Review.
-
Mycobacterial IHF is a highly dynamic nucleoid-associated protein that assists HupB in organizing chromatin.Front Microbiol. 2023 Mar 7;14:1146406. doi: 10.3389/fmicb.2023.1146406. eCollection 2023. Front Microbiol. 2023. PMID: 36960278 Free PMC article.
-
Shift and Metabolic Potentials of Microbial Eukaryotic Communities Across the Full Depths of the Mariana Trench.Front Microbiol. 2021 Jan 18;11:603692. doi: 10.3389/fmicb.2020.603692. eCollection 2020. Front Microbiol. 2021. PMID: 33537012 Free PMC article.
References
-
- Pettijohn DE. 1988. Histone-like proteins and bacterial chromosome structure. J. Biol. Chem. 263:12793–12796 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

