An integrated map of HIV-human protein complexes that facilitate viral infection
- PMID: 24817247
- PMCID: PMC4016004
- DOI: 10.1371/journal.pone.0096687
An integrated map of HIV-human protein complexes that facilitate viral infection
Abstract
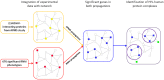
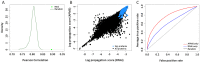
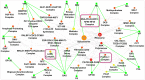
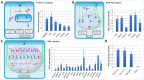
Recent proteomic and genetic studies have aimed to identify a complete network of interactions between HIV and human proteins and genes. This HIV-human interaction network provides invaluable information as to how HIV exploits the host machinery and can be used as a starting point for further functional analyses. We integrated this network with complementary datasets of protein function and interaction to nominate human protein complexes with likely roles in viral infection. Based on our approach we identified a global map of 40 HIV-human protein complexes with putative roles in HIV infection, some of which are involved in DNA replication and repair, transcription, translation, and cytoskeletal regulation. Targeted RNAi screens were used to validate several proteins and complexes for functional impact on viral infection. Thus, our HIV-human protein complex map provides a significant resource of potential HIV-host interactions for further study.
Conflict of interest statement
Figures

Similar articles
-
Global landscape of HIV-human protein complexes.Nature. 2011 Dec 21;481(7381):365-70. doi: 10.1038/nature10719. Nature. 2011. PMID: 22190034 Free PMC article.
-
Purification and characterization of HIV-human protein complexes.Methods. 2011 Jan;53(1):13-9. doi: 10.1016/j.ymeth.2010.08.007. Epub 2010 Aug 12. Methods. 2011. PMID: 20708689 Free PMC article. Review.
-
Quantitative Temporal Viromics of an Inducible HIV-1 Model Yields Insight to Global Host Targets and Phospho-Dynamics Associated with Protein Vpr.Mol Cell Proteomics. 2017 Aug;16(8):1447-1461. doi: 10.1074/mcp.M116.066019. Epub 2017 Jun 12. Mol Cell Proteomics. 2017. PMID: 28606917 Free PMC article.
-
Proteome analysis of the HIV-1 Gag interactome.Virology. 2014 Jul;460-461:194-206. doi: 10.1016/j.virol.2014.04.038. Epub 2014 Jun 10. Virology. 2014. PMID: 25010285
-
Modulation of HIV-1-host interaction: role of the Vpu accessory protein.Retrovirology. 2010 Dec 22;7:114. doi: 10.1186/1742-4690-7-114. Retrovirology. 2010. PMID: 21176220 Free PMC article. Review.
Cited by
-
Understanding Human-Virus Protein-Protein Interactions Using a Human Protein Complex-Based Analysis Framework.mSystems. 2019 Apr 9;4(2):e00303-18. doi: 10.1128/mSystems.00303-18. eCollection 2019 Mar-Apr. mSystems. 2019. PMID: 30984872 Free PMC article.
-
Characterization of HIV-1 integrase interaction with human Ku70 protein and initial implications for drug targeting.Sci Rep. 2017 Jul 17;7(1):5649. doi: 10.1038/s41598-017-05659-5. Sci Rep. 2017. PMID: 28717247 Free PMC article.
-
The Role of Ku70 as a Cytosolic DNA Sensor in Innate Immunity and Beyond.Front Cell Infect Microbiol. 2021 Oct 21;11:761983. doi: 10.3389/fcimb.2021.761983. eCollection 2021. Front Cell Infect Microbiol. 2021. PMID: 34746031 Free PMC article. Review.
-
G protein-coupled and ATP-sensitive inwardly rectifying potassium ion channels are essential for HIV entry.Sci Rep. 2019 Mar 11;9(1):4113. doi: 10.1038/s41598-019-40968-x. Sci Rep. 2019. PMID: 30858482 Free PMC article.
-
Synthesis, biological evaluation and molecular modeling of novel azaspiro dihydrotriazines as influenza virus inhibitors targeting the host factor dihydrofolate reductase (DHFR).Eur J Med Chem. 2018 Jul 15;155:229-243. doi: 10.1016/j.ejmech.2018.05.059. Epub 2018 Jun 2. Eur J Med Chem. 2018. PMID: 29886325 Free PMC article.
References
-
- Frankel AD, Young JA (1998) HIV-1: fifteen proteins and an RNA. Annu Rev Biochem 67: 1–25. - PubMed
-
- Bhattacharya S, Osman H (2009) Novel targets for anti-retroviral therapy. J Infect 59: 377–386. - PubMed
-
- Pache L, Konig R, Chanda SK (2011) Identifying HIV-1 host cell factors by genome-scale RNAi screening. Methods 53: 3–12. - PubMed
-
- Zhou H, Xu M, Huang Q, Gates AT, Zhang XD, et al. (2008) Genome-scale RNAi screen for host factors required for HIV replication. Cell Host Microbe 4: 495–504. - PubMed
-
- Brass AL, Dykxhoorn DM, Benita Y, Yan N, Engelman A, et al. (2008) Identification of host proteins required for HIV infection through a functional genomic screen. Science 319: 921–926. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

