DNA torsion as a feedback mediator of transcription and chromatin dynamics
- PMID: 24819949
- PMCID: PMC4133216
- DOI: 10.4161/nucl.29086
DNA torsion as a feedback mediator of transcription and chromatin dynamics
Abstract
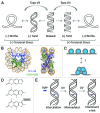
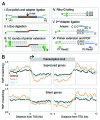
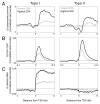
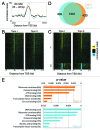
The double helical structure of DNA lends itself to topological constraints. Many DNA-based processes alter the topological state of DNA, generating torsional stress, which is efficiently relieved by topoisomerases. Maintaining this topological balance is crucial to cell survival, as excessive torsional strain risks DNA damage. Here, we review the mechanisms that generate and modulate DNA torsion within the cell. In particular, we discuss how transcription-generated torsional stress affects Pol II kinetics and chromatin dynamics, highlighting an emerging role of DNA torsion as a feedback mediator of torsion-generating processes.
Keywords: RNA Polymerase II; nascent RNA; nucleosome turnover; topoisomerase; torsional stress.
Figures
Comment on
- Teves SS, Henikoff S. Transcription-generated torsional stress destabilizes nucleosomes. Nat Struct Mol Biol. 2014;21:88–94. doi: 10.1038/nsmb.2723.
Similar articles
-
To Break or Not to Break: The Role of TOP2B in Transcription.Int J Mol Sci. 2023 Sep 30;24(19):14806. doi: 10.3390/ijms241914806. Int J Mol Sci. 2023. PMID: 37834253 Free PMC article. Review.
-
Chromatin Buffers Torsional Stress During Transcription.bioRxiv [Preprint]. 2024 Oct 18:2024.10.15.618270. doi: 10.1101/2024.10.15.618270. bioRxiv. 2024. PMID: 39464147 Free PMC article. Preprint.
-
Transcription-generated torsional stress destabilizes nucleosomes.Nat Struct Mol Biol. 2014 Jan;21(1):88-94. doi: 10.1038/nsmb.2723. Epub 2013 Dec 8. Nat Struct Mol Biol. 2014. PMID: 24317489 Free PMC article.
-
Chromatin Architectural Factors as Safeguards against Excessive Supercoiling during DNA Replication.Int J Mol Sci. 2020 Jun 24;21(12):4504. doi: 10.3390/ijms21124504. Int J Mol Sci. 2020. PMID: 32599919 Free PMC article. Review.
-
Topoisomerase II binds nucleosome-free DNA and acts redundantly with topoisomerase I to enhance recruitment of RNA Pol II in budding yeast.Proc Natl Acad Sci U S A. 2011 Aug 2;108(31):12693-8. doi: 10.1073/pnas.1106834108. Epub 2011 Jul 19. Proc Natl Acad Sci U S A. 2011. PMID: 21771901 Free PMC article.
Cited by
-
VAL1 acts as an assembly platform co-ordinating co-transcriptional repression and chromatin regulation at Arabidopsis FLC.Nat Commun. 2022 Sep 21;13(1):5542. doi: 10.1038/s41467-022-32897-7. Nat Commun. 2022. PMID: 36130923 Free PMC article.
-
Chd1 protects genome integrity at promoters to sustain hypertranscription in embryonic stem cells.Nat Commun. 2021 Aug 11;12(1):4859. doi: 10.1038/s41467-021-25088-3. Nat Commun. 2021. PMID: 34381042 Free PMC article.
-
Permanganate/S1 Nuclease Footprinting Reveals Non-B DNA Structures with Regulatory Potential across a Mammalian Genome.Cell Syst. 2017 Mar 22;4(3):344-356.e7. doi: 10.1016/j.cels.2017.01.013. Epub 2017 Feb 22. Cell Syst. 2017. PMID: 28237796 Free PMC article.
-
Downstream-of-gene (DoG) transcripts contribute to an imbalance in the cancer cell transcriptome.Sci Adv. 2024 Jul 5;10(27):eadh9613. doi: 10.1126/sciadv.adh9613. Epub 2024 Jul 3. Sci Adv. 2024. PMID: 38959318 Free PMC article.
-
Protein/DNA interactions in complex DNA topologies: expect the unexpected.Biophys Rev. 2016;8(3):233-243. doi: 10.1007/s12551-016-0208-8. Epub 2016 Aug 8. Biophys Rev. 2016. PMID: 27738452 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
