De novo transcriptome assembly and analyses of gene expression during photomorphogenesis in diploid wheat Triticum monococcum
- PMID: 24821410
- PMCID: PMC4018402
- DOI: 10.1371/journal.pone.0096855
De novo transcriptome assembly and analyses of gene expression during photomorphogenesis in diploid wheat Triticum monococcum
Erratum in
- PLoS One. 2014;9(8):e105275
Abstract
Background: Triticum monococcum (2n) is a close ancestor of T. urartu, the A-genome progenitor of cultivated hexaploid wheat, and is therefore a useful model for the study of components regulating photomorphogenesis in diploid wheat. In order to develop genetic and genomic resources for such a study, we constructed genome-wide transcriptomes of two Triticum monococcum subspecies, the wild winter wheat T. monococcum ssp. aegilopoides (accession G3116) and the domesticated spring wheat T. monococcum ssp. monococcum (accession DV92) by generating de novo assemblies of RNA-Seq data derived from both etiolated and green seedlings.
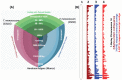
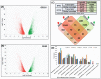
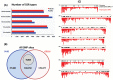
Principal findings: The de novo transcriptome assemblies of DV92 and G3116 represent 120,911 and 117,969 transcripts, respectively. We successfully mapped ∼90% of these transcripts from each accession to barley and ∼95% of the transcripts to T. urartu genomes. However, only ∼77% transcripts mapped to the annotated barley genes and ∼85% transcripts mapped to the annotated T. urartu genes. Differential gene expression analyses revealed 22% more light up-regulated and 35% more light down-regulated transcripts in the G3116 transcriptome compared to DV92. The DV92 and G3116 mRNA sequence reads aligned against the reference barley genome led to the identification of ∼500,000 single nucleotide polymorphism (SNP) and ∼22,000 simple sequence repeat (SSR) sites.
Conclusions: De novo transcriptome assemblies of two accessions of the diploid wheat T. monococcum provide new empirical transcriptome references for improving Triticeae genome annotations, and insights into transcriptional programming during photomorphogenesis. The SNP and SSR sites identified in our analysis provide additional resources for the development of molecular markers.
Conflict of interest statement
Figures
Similar articles
-
Genome-wide polymorphisms from RNA sequencing assembly of leaf transcripts facilitate phylogenetic analysis and molecular marker development in wild einkorn wheat.Mol Genet Genomics. 2019 Oct;294(5):1327-1341. doi: 10.1007/s00438-019-01581-9. Epub 2019 Jun 11. Mol Genet Genomics. 2019. PMID: 31187273
-
Molecular characterisation of two novel starch granule proteins 1 in wild and cultivated diploid A genome wheat species.J Plant Res. 2018 May;131(3):487-496. doi: 10.1007/s10265-017-1005-6. Epub 2017 Dec 19. J Plant Res. 2018. PMID: 29260339
-
An integrated molecular linkage map of diploid wheat based on a Triticum boeoticum x T. monococcum RIL population.Theor Appl Genet. 2007 Aug;115(3):301-12. doi: 10.1007/s00122-007-0543-z. Epub 2007 May 25. Theor Appl Genet. 2007. PMID: 17565482
-
Development of a New Am -Genome-Specific Single Nucleotide Polymorphism Marker Set for the Molecular Characterization of Wheat-Triticum monococcum Introgression Lines.Plant Genome. 2019 Nov;12(3):1-7. doi: 10.3835/plantgenome2018.12.0098. Plant Genome. 2019. PMID: 33016586
-
RNA-Seq-based DNA marker analysis of the genetics and molecular evolution of Triticeae species.Funct Integr Genomics. 2021 Nov;21(5-6):535-542. doi: 10.1007/s10142-021-00799-4. Epub 2021 Aug 18. Funct Integr Genomics. 2021. PMID: 34405283 Review.
Cited by
-
FISH karyotype comparison between Ab- and A-genome chromosomes using oligonucleotide probes.J Appl Genet. 2020 Sep;61(3):313-322. doi: 10.1007/s13353-020-00555-7. Epub 2020 Apr 5. J Appl Genet. 2020. PMID: 32248406
-
Identification and characterization of wheat stem rust resistance gene Sr21 effective against the Ug99 race group at high temperature.PLoS Genet. 2018 Apr 3;14(4):e1007287. doi: 10.1371/journal.pgen.1007287. eCollection 2018 Apr. PLoS Genet. 2018. PMID: 29614079 Free PMC article.
-
Diploid genome differentiation conferred by RNA sequencing-based survey of genome-wide polymorphisms throughout homoeologous loci in Triticum and Aegilops.BMC Genomics. 2020 Mar 20;21(1):246. doi: 10.1186/s12864-020-6664-3. BMC Genomics. 2020. PMID: 32192452 Free PMC article.
-
Insights into Four NAC Transcription Factors Involved in Grain Development and in Response to Moderate Heat in the Triticeae Tribe.Int J Mol Sci. 2022 Oct 2;23(19):11672. doi: 10.3390/ijms231911672. Int J Mol Sci. 2022. PMID: 36232974 Free PMC article.
-
RNA-seq analysis reveals considerable genetic diversity and provides genetic markers saturating all chromosomes in the diploid wild wheat relative Aegilops umbellulata.BMC Plant Biol. 2018 Nov 8;18(1):271. doi: 10.1186/s12870-018-1498-8. BMC Plant Biol. 2018. PMID: 30409135 Free PMC article.
References
-
- Heun M, Schäfer-Pregl R, Klawan D, Castagna R, Accerbi M, et al. (1997) Site of Einkorn Wheat Domestication Identified by DNA Fingerprinting. Science 278: 1312–1314 10.1126/science.278.5341.1312 - DOI
-
- Bennett MD, Leitch IJ (1995) Nuclear DNA Amounts in Angiosperms. Ann Bot 76: 113–176 10.1006/anbo.1995.1085 - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

