Identification of Horizontally-transferred Genomic Islands and Genome Segmentation Points by Using the GC Profile Method
- PMID: 24822029
- PMCID: PMC4009839
- DOI: 10.2174/1389202915999140328163125
Identification of Horizontally-transferred Genomic Islands and Genome Segmentation Points by Using the GC Profile Method
Abstract
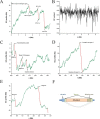
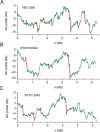
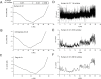
The nucleotide composition of genomes undergoes dramatic variations among all three kingdoms of life. GC content, an important characteristic for a genome, is related to many important functions, and therefore GC content and its distribution are routinely reported for sequenced genomes. Traditionally, GC content distribution is assessed by computing GC contents in windows that slide along the genome. Disadvantages of this routinely used window-based method include low resolution and low sensitivity. Additionally, different window sizes result in different GC content distribution patterns within the same genome. We proposed a windowless method, the GC profile, for displaying GC content variations across the genome. Compared to the window-based method, the GC profile has the following advantages: 1) higher sensitivity, because of variation-amplifying procedures; 2) higher resolution, because boundaries between domains can be determined at one single base pair; 3) uniqueness, because the GC profile is unique for a given genome and 4) the capacity to show both global and regional GC content distributions. These characteristics are useful in identifying horizontally-transferred genomic islands and homogenous GC-content domains. Here, we review the applications of the GC profile in identifying genomic islands and genome segmentation points, and in serving as a platform to integrate with other algorithms for genome analysis. A web server generating GC profiles and implementing relevant genome segmentation algorithms is available at: www.zcurve.net.
Keywords: GC profile; Genome segmentation.; Genomic island.
Figures

Similar articles
-
A systematic method to identify genomic islands and its applications in analyzing the genomes of Corynebacterium glutamicum and Vibrio vulnificus CMCP6 chromosome I.Bioinformatics. 2004 Mar 22;20(5):612-22. doi: 10.1093/bioinformatics/btg453. Epub 2004 Jan 22. Bioinformatics. 2004. PMID: 15033867
-
GC-Profile: a web-based tool for visualizing and analyzing the variation of GC content in genomic sequences.Nucleic Acids Res. 2006 Jul 1;34(Web Server issue):W686-91. doi: 10.1093/nar/gkl040. Nucleic Acids Res. 2006. PMID: 16845098 Free PMC article.
-
Isochore structures in the chicken genome.FEBS J. 2006 Apr;273(8):1637-48. doi: 10.1111/j.1742-4658.2006.05178.x. FEBS J. 2006. PMID: 16623701
-
Investigating genomic structure using changept: A Bayesian segmentation model.Comput Struct Biotechnol J. 2014 Aug 27;10(17):107-15. doi: 10.1016/j.csbj.2014.08.003. eCollection 2014 Jul. Comput Struct Biotechnol J. 2014. PMID: 25349679 Free PMC article. Review.
-
Sequence segmentation.Methods Mol Biol. 2008;452:207-29. doi: 10.1007/978-1-60327-159-2_11. Methods Mol Biol. 2008. PMID: 18566767 Review.
Cited by
-
Genomic Analysis of Pasteurella atlantica Provides Insight on Its Virulence Factors and Phylogeny and Highlights the Potential of Reverse Vaccinology in Aquaculture.Microorganisms. 2021 Jun 4;9(6):1215. doi: 10.3390/microorganisms9061215. Microorganisms. 2021. PMID: 34199775 Free PMC article.
-
Identification and analysis of genomic islands in Burkholderia cenocepacia AU 1054 with emphasis on pathogenicity islands.BMC Microbiol. 2017 Mar 27;17(1):73. doi: 10.1186/s12866-017-0986-6. BMC Microbiol. 2017. PMID: 28347342 Free PMC article.
-
Zisland Explorer: detect genomic islands by combining homogeneity and heterogeneity properties.Brief Bioinform. 2017 May 1;18(3):357-366. doi: 10.1093/bib/bbw019. Brief Bioinform. 2017. PMID: 26992782 Free PMC article.
-
Computational methods for predicting genomic islands in microbial genomes.Comput Struct Biotechnol J. 2016 May 7;14:200-6. doi: 10.1016/j.csbj.2016.05.001. eCollection 2016. Comput Struct Biotechnol J. 2016. PMID: 27293536 Free PMC article. Review.
-
Colonization, Infection, and the Accessory Genome of Klebsiella pneumoniae.Front Cell Infect Microbiol. 2018 Jan 22;8:4. doi: 10.3389/fcimb.2018.00004. eCollection 2018. Front Cell Infect Microbiol. 2018. PMID: 29404282 Free PMC article. Review.
References
-
- Nelson KE, Clayton RA, Gill SR, Gwinn ML, Dodson RJ, Haft DH, Hickey EK, Peterson JD, Nelson WC, Ketchum KA, McDonald L, Utterback TR, Malek JA, Linher KD, Garrett MM, Stewart AM, Cotton MD, Pratt MS, Phillips CA, Richardson D, Heidelberg J, Sutton GG, Fleischmann RD, Eisen JA, White O, Salzberg SL, Smith HO, Venter JC, Fraser CM. Evidence for lateral gene transfer between Archaea and bacteria from genome sequence of Thermotoga maritima. Nature. 1999;399(6734):323–329. - PubMed
-
- Ochman H. Lateral and oblique gene transfer. Curr. Opin. Genet. Dev. 2001;11(6):616–619. - PubMed
-
- Ochman H, Lawrence JG, Groisman EA. Lateral gene transfer and the nature of bacterial innovation. Nature. 2000;405(6784):299–304. - PubMed
-
- Hacker J, Kaper JB. Pathogenicity islands and the evolution of microbes. Annu Rev Microbiol. 2000;54:641–679. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous
