Mitochondrial genome of Phlebia radiata is the second largest (156 kbp) among fungi and features signs of genome flexibility and recent recombination events
- PMID: 24824642
- PMCID: PMC4019555
- DOI: 10.1371/journal.pone.0097141
Mitochondrial genome of Phlebia radiata is the second largest (156 kbp) among fungi and features signs of genome flexibility and recent recombination events
Abstract
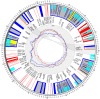
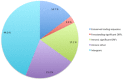
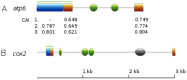
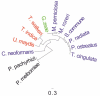
Mitochondria are eukaryotic organelles supporting individual life-style via generation of proton motive force and cellular energy, and indispensable metabolic pathways. As part of genome sequencing of the white rot Basidiomycota species Phlebia radiata, we first assembled its mitochondrial genome (mtDNA). So far, the 156 348 bp mtDNA is the second largest described for fungi, and of considerable size among eukaryotes. The P. radiata mtDNA assembled as single circular dsDNA molecule containing genes for the large and small ribosomal RNAs, 28 transfer RNAs, and over 100 open reading frames encoding the 14 fungal conserved protein subunits of the mitochondrial complexes I, III, IV, and V. Two genes (atp6 and tRNA-IleGAU) were duplicated within 6.1 kbp inverted region, which is a unique feature of the genome. The large mtDNA size, however, is explained by the dominance of intronic and intergenic regions (sum 80% of mtDNA sequence). The intergenic DNA stretches harness short (≤ 200 nt) repetitive, dispersed and overlapping sequence elements in abundance. Long self-splicing introns of types I and II interrupt eleven of the conserved genes (cox1,2,3; cob; nad1,2,4,4L,5; rnl; rns). The introns embrace a total of 57 homing endonucleases with LAGLIDADGD and GYI-YIG core motifs, which makes P. radiata mtDNA to one of the largest known reservoirs of intron-homing endonucleases. The inverted duplication, intergenic stretches, and intronic features are indications of dynamics and genetic flexibility of the mtDNA, not fully recognized to this extent in fungal mitochondrial genomes previously, thus giving new insights for the evolution of organelle genomes in eukaryotes.
Conflict of interest statement
Figures


Similar articles
-
Annotation and analysis of the mitochondrial genome of Coniothyrium glycines, causal agent of red leaf blotch of soybean, reveals an abundance of homing endonucleases.PLoS One. 2018 Nov 7;13(11):e0207062. doi: 10.1371/journal.pone.0207062. eCollection 2018. PLoS One. 2018. PMID: 30403741 Free PMC article.
-
Sequencing of mitochondrial genomes of nine Aspergillus and Penicillium species identifies mobile introns and accessory genes as main sources of genome size variability.BMC Genomics. 2012 Dec 12;13:698. doi: 10.1186/1471-2164-13-698. BMC Genomics. 2012. PMID: 23234273 Free PMC article.
-
The mitochondrial genome of Morchella importuna (272.2 kb) is the largest among fungi and contains numerous introns, mitochondrial non-conserved open reading frames and repetitive sequences.Int J Biol Macromol. 2020 Jan 15;143:373-381. doi: 10.1016/j.ijbiomac.2019.12.056. Epub 2019 Dec 9. Int J Biol Macromol. 2020. PMID: 31830457
-
Evolution and maintenance of mtDNA gene content across eukaryotes.Biochem J. 2024 Aug 7;481(15):1015-1042. doi: 10.1042/BCJ20230415. Biochem J. 2024. PMID: 39101615 Free PMC article. Review.
-
Insights into Fungal Mitochondrial Genomes and Inheritance Based on Current Findings from Yeast-like Fungi.J Fungi (Basel). 2024 Jun 21;10(7):441. doi: 10.3390/jof10070441. J Fungi (Basel). 2024. PMID: 39057326 Free PMC article. Review.
Cited by
-
The First Mitochondrial Genome for Geastrales (Sphaerobolus stellatus) Reveals Intron Dynamics and Large-Scale Gene Rearrangements of Basidiomycota.Front Microbiol. 2020 Aug 11;11:1970. doi: 10.3389/fmicb.2020.01970. eCollection 2020. Front Microbiol. 2020. PMID: 32849488 Free PMC article.
-
Lignocellulose-converting enzyme activity profiles correlate with molecular systematics and phylogeny grouping in the incoherent genus Phlebia (Polyporales, Basidiomycota).BMC Microbiol. 2015 Oct 19;15:217. doi: 10.1186/s12866-015-0538-x. BMC Microbiol. 2015. PMID: 26482661 Free PMC article.
-
Analysis of the Mitochondrial Genome in Hypomyces aurantius Reveals a Novel Twintron Complex in Fungi.Int J Mol Sci. 2016 Jun 30;17(7):1049. doi: 10.3390/ijms17071049. Int J Mol Sci. 2016. PMID: 27376282 Free PMC article.
-
Comparative Mitochondrial Genome Analysis of Two Ectomycorrhizal Fungi (Rhizopogon) Reveals Dynamic Changes of Intron and Phylogenetic Relationships of the Subphylum Agaricomycotina.Int J Mol Sci. 2019 Oct 18;20(20):5167. doi: 10.3390/ijms20205167. Int J Mol Sci. 2019. PMID: 31635252 Free PMC article.
-
Comparative mitogenome analysis of two ectomycorrhizal fungi (Paxillus) reveals gene rearrangement, intron dynamics, and phylogeny of basidiomycetes.IMA Fungus. 2020 Jul 2;11:12. doi: 10.1186/s43008-020-00038-8. eCollection 2020. IMA Fungus. 2020. PMID: 32670777 Free PMC article.
References
-
- Nakasone KK, Sytsma KJ (1993) Biosystematic studies on Phlebia acerina, P. rufa, and P. radiata in North America. Mycologia 85: 996–1016.
-
- Miettinen O, Larsson E, Sjökvist E, Larsson KH (2012) Comprehensive taxon sampling reveals unaccounted diversity and morphological plasticity in a group of dimitic polypores (Polyporales, Basidiomycota). Cladistics 28: 251–270. - PubMed
-
- Moilanen AM, Lundell T, Vares T, Hatakka A (1996) Manganese and malonate are individual regulators for the production of lignin and manganese peroxidase isozymes and in the degradation of lignin by Phlebia radiata . Appl Microbiol Biotechnol 45: 792–799.
-
- Lundell TK, Mäkelä MR, Hildén K (2010) Lignin-modifying enzymes in filamentous basidiomycetes - ecological, functional and phylogenetic review. J Basic Microbiol 50: 5–20. - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

