Wnt signaling directs a metabolic program of glycolysis and angiogenesis in colon cancer
- PMID: 24825347
- PMCID: PMC4194089
- DOI: 10.15252/embj.201488598
Wnt signaling directs a metabolic program of glycolysis and angiogenesis in colon cancer
Abstract
Much of the mechanism by which Wnt signaling drives proliferation during oncogenesis is attributed to its regulation of the cell cycle. Here, we show how Wnt/β-catenin signaling directs another hallmark of tumorigenesis, namely Warburg metabolism. Using biochemical assays and fluorescence lifetime imaging microscopy (FLIM) to probe metabolism in vitro and in living tumors, we observe that interference with Wnt signaling in colon cancer cells reduces glycolytic metabolism and results in small, poorly perfused tumors. We identify pyruvate dehydrogenase kinase 1 (PDK1) as an important direct target within a larger gene program for metabolism. PDK1 inhibits pyruvate flux to mitochondrial respiration and a rescue of its expression in Wnt-inhibited cancer cells rescues glycolysis as well as vessel growth in the tumor microenvironment. Thus, we identify an important mechanism by which Wnt-driven Warburg metabolism directs the use of glucose for cancer cell proliferation and links it to vessel delivery of oxygen and nutrients.
Keywords: Wnt; angiogenesis; colon cancer; fluorescence lifetime imaging; metabolism.
© 2014 The Authors.
Figures
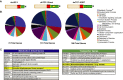
Schematic of dnLEF/TCF isoforms expressed (beige box = context-dependent regulatory domain; red box = HMG DNA binding domain; yellow box = nuclear localization signal; green box = alternatively spliced C-terminal tails; yellow star = mutation in E-tail).
Ontology analysis of dnLEF/TCF-downregulated genes reveals a large category of genes linked to metabolism. PANTHER binomial statistical analysis determined statistically significant overrepresentation of regulated genes within each category compared to representation in the human genome (*P-value < 0.01).
List of metabolism-linked genes downregulated by all three dnLEF/TCF isoforms. List of transporter genes regulated by at least 2 dnLEF/TCF isoforms. For all genes on the list changes are P < 0.05, and range of fold changes is −1.3 to −6.0.

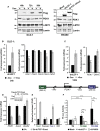
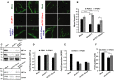
Western blot analysis of lysates collected from DLD-1 dnLEF-1(2) cells treated with or without 0.01 μg/ml doxycycline for 24 h and with or without PDK1 lentivirus for 72 h. FLIM imaging was performed at confluency (96 h doxycycline and 7 days post-transduction). Intensity images are shown on the left. FLIM results, shown through the free/bound NADH color mapping (right), as well as the scatterplot, show that doxycycline induction of dnLEF-1 shifted the phasor toward bound NADH, while PDK1 rescue shifted it back to its original position (P < 0.0001 comparing −Dox to +Dox and comparing +Dox to +Dox+PDK1).
FLIM analysis of SW480 cells treated with 50 mM DCA for 48 h shows a phasor shift toward bound NADH (P < 0.0001 comparing +DCA to mock).
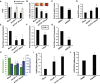
OCR/ECAR ratio of DCA (10 mM)-treated SW480 cells shows that blocking PDK1 activity doubles the oxygen consumption rate (mitochondrial activity) relative to the extracellular acidification rate (glycolysis-produced lactate). Data shown represent the average of three independent trials (± SD; *P-value < 0.05).
Sulforhodamine B cell proliferation assay of SW480 cells treated with or without 2.5 μM irinotecan, 20 mM DCA, and dnLEF-1 lentivirus shows an increased sensitivity to irinotecan when treated with DCA or dnLEF-1. A representative graph of two trials is shown. Error bars represent the standard deviation between eight internal replicates.
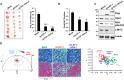
Expression of dnLEF-1 or dnTCF-1Emut in SW480 cells results in smaller xenograft tumors. Images of tumors are shown with quantification of the tumor mass and volume at the time of harvest. Data include measurements of eight tumors for each condition. Error bars represent the SEM among eight replicates (***P-value < 0.001).
Ki67 staining of paraffin-embedded sections from xenograft tumors shows fewer Ki67-positive cells with dnLEF-1/dnTCF-Emut expression. Data shown represent the average of counts from at least eight fields. Error bars represent the SEM among at least eight replicates (*P-value < 0.05).
Western blot was performed on protein lysates prepared from freshly extracted xenograft tumors. Western blot analysis of endogenous PDK1 and its target, pyruvate dehydrogenase (pSer293-PDH), shows decreases when dnLEF-1 or dnTCF-1Emut tumors are expressed. cMYC expression shows variable levels of expression.
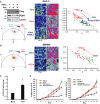
Phasor plot representation for the color mapping of in vivo tumor FLIM analysis, fluorescence intensity (top image panels), FLIM color mapping (bottom panels), and scatterplot analysis are as described in Fig5 except that these analyses were performed on living, surgically exposed, yet still actively perfused, xenograft tumors. Both the free/bound NADH color mapping (bottom row) and scatterplot (average phasor position of individual cells within each tumor) show a shift in the phasor position toward bound NADH with dnLEF-1 expression and a return to free NADH with PDK1 overexpression (P < 0.0001 comparing dnLEF-1 to mock and comparing dnLEF-1+PDK1 to dnLEF-1). A minimum of three fields of view per tumor were analyzed. Data shown are from one mouse representative of eight replicates (additional tumor analysis in Supplementary Fig S7).

Model of Wnt regulation of cancer metabolism and angiogenesis through PDK1.
Immunohistochemical staining of normal human small intestine (PDK1) and normal human colon (phospho-PDH and HIF1α) shows correlations between high levels of Wnt signaling, PDK1 activity, and HIF1α levels. Bottom row shows higher-power images of boxed portions in the top row.
Comment in
-
Wnt meets Warburg: another piece in the puzzle?EMBO J. 2014 Jul 1;33(13):1420-2. doi: 10.15252/embj.201488785. Epub 2014 May 19. EMBO J. 2014. PMID: 24843042 Free PMC article.
References
-
- Baumunk D, Reichelt U, Hildebrandt J, Krause H, Ebbing J, Cash H, Miller K, Schostak M, Weikert S. Expression parameters of the metabolic pathway genes pyruvate dehydrogenase kinase-1 (PDK-1) and DJ-1/PARK7 in renal cell carcinoma (RCC) World J Urol. 2012;31:1191–1196. - PubMed
-
- Becker W, Bergmann A, Hink MA, König K, Benndorf K, Biskup C. Fluorescence lifetime imaging by time-correlated single-photon counting. Microsc Res Tech. 2004;63:58–66. - PubMed
-
- Bienz M, Clevers H. Linking colorectal cancer to Wnt signaling review. Cell. 2000;103:311–320. - PubMed
-
- Bird DK, Yan L, Vrotsos KM, Eliceiri KW, Vaughan EM, Keely PJ, White JG, Ramanujam N. Metabolic mapping of MCF10A human breast cells via multiphoton fluorescence lifetime imaging of the coenzyme NADH. Cancer Res. 2005;65:8766–8773. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA185189/CA/NCI NIH HHS/United States
- R01 CA090571/CA/NCI NIH HHS/United States
- U01 AA021838/AA/NIAAA NIH HHS/United States
- T32 GM007185/GM/NIGMS NIH HHS/United States
- CA108697/CA/NCI NIH HHS/United States
- CA-62203/CA/NCI NIH HHS/United States
- P50 GM076516/GM/NIGMS NIH HHS/United States
- P41-RR03155/RR/NCRR NIH HHS/United States
- T32GM007185/GM/NIGMS NIH HHS/United States
- P01GM081621/GM/NIGMS NIH HHS/United States
- P30CA062203/CA/NCI NIH HHS/United States
- R01 GM073981/GM/NIGMS NIH HHS/United States
- R01CA156674/CA/NCI NIH HHS/United States
- R01 CA096878/CA/NCI NIH HHS/United States
- CA096878/CA/NCI NIH HHS/United States
- R01 CA156674/CA/NCI NIH HHS/United States
- R01CA90571/CA/NCI NIH HHS/United States
- R01 CA108697/CA/NCI NIH HHS/United States
- P41 GM103540/GM/NIGMS NIH HHS/United States
- P30 CA062203/CA/NCI NIH HHS/United States
- R01CA185189/CA/NCI NIH HHS/United States
- P01 GM081621/GM/NIGMS NIH HHS/United States
- R01GM073981/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

