Evidence for increased levels of positive and negative selection on the X chromosome versus autosomes in humans
- PMID: 24830675
- PMCID: PMC4137703
- DOI: 10.1093/molbev/msu166
Evidence for increased levels of positive and negative selection on the X chromosome versus autosomes in humans
Abstract
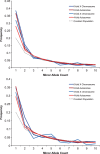
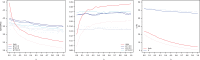
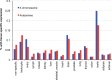
Partially recessive variants under positive selection are expected to go to fixation more quickly on the X chromosome as a result of hemizygosity, an effect known as faster-X. Conversely, purifying selection is expected to reduce substitution rates more effectively on the X chromosome. Previous work in humans contrasted divergence on the autosomes and X chromosome, with results tending to support the faster-X effect. However, no study has yet incorporated both divergence and polymorphism to quantify the effects of both purifying and positive selection, which are opposing forces with respect to divergence. In this study, we develop a framework that integrates previously developed theory addressing differential rates of X and autosomal evolution with methods that jointly estimate the level of purifying and positive selection via modeling of the distribution of fitness effects (DFE). We then utilize this framework to estimate the proportion of nonsynonymous substitutions fixed by positive selection (α) using exome sequence data from a West African population. We find that varying the female to male breeding ratio (β) has minimal impact on the DFE for the X chromosome, especially when compared with the effect of varying the dominance coefficient of deleterious alleles (h). Estimates of α range from 46% to 51% and from 4% to 24% for the X chromosome and autosomes, respectively. While dependent on h, the magnitude of the difference between α values estimated for these two systems is highly statistically significant over a range of biologically realistic parameter values, suggesting faster-X has been operating in humans.
Keywords: X chromosome; autosomes; distribution of fitness effects; humans; positive selection; purifying selection.
© The Author 2014. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution. All rights reserved. For permissions, please e-mail: journals.permissions@oup.com.
Figures
Similar articles
-
Hemizygosity Enhances Purifying Selection: Lack of Fast-Z Evolution in Two Satyrine Butterflies.Genome Biol Evol. 2016 Oct 23;8(10):3108-3119. doi: 10.1093/gbe/evw214. Genome Biol Evol. 2016. PMID: 27590089 Free PMC article.
-
Sex differences in deleterious mutational effects in Drosophila melanogaster: combining quantitative and population genetic insights.Genetics. 2021 Nov 5;219(3):iyab143. doi: 10.1093/genetics/iyab143. Genetics. 2021. PMID: 34740242 Free PMC article.
-
Evidence for widespread positive and purifying selection across the European rabbit (Oryctolagus cuniculus) genome.Mol Biol Evol. 2012 Jul;29(7):1837-49. doi: 10.1093/molbev/mss025. Epub 2012 Jan 31. Mol Biol Evol. 2012. PMID: 22319161 Free PMC article.
-
Evolution on the X chromosome: unusual patterns and processes.Nat Rev Genet. 2006 Aug;7(8):645-53. doi: 10.1038/nrg1914. Nat Rev Genet. 2006. PMID: 16847464 Review.
-
Effects of new mutations on fitness: insights from models and data.Ann N Y Acad Sci. 2014 Jul;1320(1):76-92. doi: 10.1111/nyas.12460. Epub 2014 May 30. Ann N Y Acad Sci. 2014. PMID: 24891070 Free PMC article. Review.
Cited by
-
Constraining models of dominance for nonsynonymous mutations in the human genome.PLoS Genet. 2024 Sep 20;20(9):e1011198. doi: 10.1371/journal.pgen.1011198. eCollection 2024 Sep. PLoS Genet. 2024. PMID: 39302992 Free PMC article.
-
Strong Selective Sweeps on the X Chromosome in the Human-Chimpanzee Ancestor Explain Its Low Divergence.PLoS Genet. 2015 Aug 14;11(8):e1005451. doi: 10.1371/journal.pgen.1005451. eCollection 2015 Aug. PLoS Genet. 2015. PMID: 26274919 Free PMC article.
-
Ancient gene flow from early modern humans into Eastern Neanderthals.Nature. 2016 Feb 25;530(7591):429-33. doi: 10.1038/nature16544. Epub 2016 Feb 17. Nature. 2016. PMID: 26886800 Free PMC article.
-
A Comparison of Selective Pressures in Plant X-Linked and Autosomal Genes.Genes (Basel). 2018 May 3;9(5):234. doi: 10.3390/genes9050234. Genes (Basel). 2018. PMID: 29751495 Free PMC article.
-
Adaptive Evolution Patterns in the Pacific Oyster Crassostrea gigas.Mar Biotechnol (NY). 2019 Oct;21(5):614-622. doi: 10.1007/s10126-019-09906-w. Epub 2019 Jun 15. Mar Biotechnol (NY). 2019. PMID: 31203476
References
-
- Asmann YW, Necela BM, Kalari KR, Hossain A, Baker TR, Carr JM, Davis C, Getz JE, Hostetter G, Li X, et al. Detection of redundant fusion transcripts as biomarkers or disease-specific therapeutic targets in breast cancer. Cancer Res. 2012;72:1921–1928. - PubMed
-
- Charlesworth B, Coyne JA, Barton NH. The relative rates of evolution of sex chromosomes and autosomes. Am Nat. 1987;130:113–146.
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

