Revised Backbone-Virtual-Bond-Angle Potentials to Treat the l- and d-Amino Acid Residues in the Coarse-Grained United Residue (UNRES) Force Field
- PMID: 24839411
- PMCID: PMC4020588
- DOI: 10.1021/ct500119r
Revised Backbone-Virtual-Bond-Angle Potentials to Treat the l- and d-Amino Acid Residues in the Coarse-Grained United Residue (UNRES) Force Field
Abstract
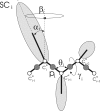
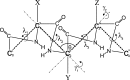
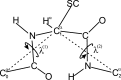
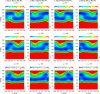
Continuing our effort to introduce d-amino-acid residues in the united residue (UNRES) force field developed in our laboratory, in this work the Cα ··· Cα ··· Cα backbone-virtual-bond-valence-angle (θ) potentials for systems containing d-amino-acid residues have been developed. The potentials were determined by integrating the combined energy surfaces of all possible triplets of terminally blocked glycine, alanine, and proline obtained with ab initio molecular quantum mechanics at the MP2/6-31G(d,p) level to calculate the corresponding potentials of mean force (PMFs). Subsequently, analytical expressions were fitted to the PMFs to give the virtual-bond-valence potentials to be used in UNRES. Alanine represented all types of amino-acid residues except glycine and proline. The blocking groups were either the N-acetyl and N',N'-dimethyl or N-acetyl and pyrrolidyl group, depending on whether the residue next in sequence was an alanine-type or a proline residue. A total of 126 potentials (63 symmetry-unrelated potentials for each set of terminally blocking groups) were determined. Together with the torsional, double-torsional, and side-chain-rotamer potentials for polypeptide chains containing d-amino-acid residues determined in our earlier work (Sieradzan et al. J. Chem. Theory Comput., 2012, 8, 4746), the new virtual-bond-angle (θ) potentials now constitute the complete set of physics-based potentials with which to run coarse-grained simulations of systems containing d-amino-acid residues. The ability of the extended UNRES force field to reproduce thermodynamics of polypeptide systems with d-amino-acid residues was tested by comparing the experimentally measured and the calculated free energies of helix formation of model KLALKLALxxLKLALKLA peptides, where x denotes any d- or l- amino-acid residue. The obtained results demonstrate that the UNRES force field with the new potentials reproduce the changes of free energies of helix formation upon d-substitution but overestimate the free energies of helix formation. To test the ability of UNRES with the new potentials to reproduce the structures of polypeptides with d-amino-acid residues, an ab initio replica-exchange folding simulation of thurincin H from Bacillus thuringiensis, which has d-amino-acid residues in the sequence, was carried out. UNRES was able to locate the native α-helical hairpin structure as the dominant structure even though no native sulfide-carbon bonds were present in the simulation.
Figures
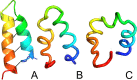
Similar articles
-
Physics-based potentials for the coupling between backbone- and side-chain-local conformational states in the UNited RESidue (UNRES) force field for protein simulations.J Chem Theory Comput. 2015 Feb 10;11(2):817-31. doi: 10.1021/ct500736a. J Chem Theory Comput. 2015. PMID: 25691834 Free PMC article.
-
Extension of UNRES force field to treat polypeptide chains with D-amino-acid residues.J Chem Theory Comput. 2012 Nov 13;8(11):4746-4757. doi: 10.1021/ct3005563. J Chem Theory Comput. 2012. PMID: 24729761 Free PMC article.
-
Determination of side-chain-rotamer and side-chain and backbone virtual-bond-stretching potentials of mean force from AM1 energy surfaces of terminally-blocked amino-acid residues, for coarse-grained simulations of protein structure and folding. II. Results, comparison with statistical potentials, and implementation in the UNRES force field.J Comput Chem. 2010 Apr 30;31(6):1154-67. doi: 10.1002/jcc.21402. J Comput Chem. 2010. PMID: 20017135 Free PMC article.
-
Coarse-grained force field: general folding theory.Phys Chem Chem Phys. 2011 Oct 14;13(38):16890-901. doi: 10.1039/c1cp20752k. Epub 2011 Jun 3. Phys Chem Chem Phys. 2011. PMID: 21643583 Free PMC article. Review.
-
Toward Consistent Physics-Based Modeling of Local Backbone Structures and Chirality Change of Proteins in Coarse-Grained Approaches.J Phys Chem Lett. 2023 Nov 9;14(44):9824-9833. doi: 10.1021/acs.jpclett.3c01988. Epub 2023 Oct 27. J Phys Chem Lett. 2023. PMID: 37889895 Free PMC article. Review.
Cited by
-
Studies of conformational changes of an arginine-binding protein from Thermotoga maritima in the presence and absence of ligand via molecular dynamics simulations with the coarse-grained UNRES force field.J Mol Model. 2015 Mar;21(3):64. doi: 10.1007/s00894-015-2609-1. Epub 2015 Mar 3. J Mol Model. 2015. PMID: 25731877
-
Physics-based potentials for the coupling between backbone- and side-chain-local conformational states in the UNited RESidue (UNRES) force field for protein simulations.J Chem Theory Comput. 2015 Feb 10;11(2):817-31. doi: 10.1021/ct500736a. J Chem Theory Comput. 2015. PMID: 25691834 Free PMC article.
-
Modeling the Structure, Dynamics, and Transformations of Proteins with the UNRES Force Field.Methods Mol Biol. 2022;2376:399-416. doi: 10.1007/978-1-0716-1716-8_23. Methods Mol Biol. 2022. PMID: 34845623
-
Residue-Level Contact Reveals Modular Domain Interactions of PICK1 Are Driven by Both Electrostatic and Hydrophobic Forces.Front Mol Biosci. 2021 Jan 27;7:616135. doi: 10.3389/fmolb.2020.616135. eCollection 2020. Front Mol Biosci. 2021. PMID: 33585564 Free PMC article.
-
When Order Meets Disorder: Modeling and Function of the Protein Interface in Fuzzy Complexes.Biomolecules. 2021 Oct 16;11(10):1529. doi: 10.3390/biom11101529. Biomolecules. 2021. PMID: 34680162 Free PMC article. Review.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
