The PIKE homolog Centaurin gamma regulates developmental timing in Drosophila
- PMID: 24845618
- PMCID: PMC4028201
- DOI: 10.1371/journal.pone.0097332
The PIKE homolog Centaurin gamma regulates developmental timing in Drosophila
Abstract
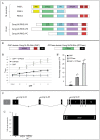
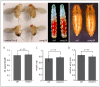
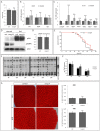
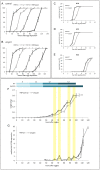
Phosphoinositide-3-kinase enhancer (PIKE) proteins encoded by the PIKE/CENTG1 gene are members of the gamma subgroup of the Centaurin superfamily of small GTPases. They are characterized by their chimeric protein domain architecture consisting of a pleckstrin homology (PH) domain, a GTPase-activating (GAP) domain, Ankyrin repeats as well as an intrinsic GTPase domain. In mammals, three PIKE isoforms with variations in protein structure and subcellular localization are encoded by the PIKE locus. PIKE inactivation in mice results in a broad range of defects, including neuronal cell death during brain development and misregulation of mammary gland development. PIKE -/- mutant mice are smaller, contain less white adipose tissue, and show insulin resistance due to misregulation of AMP-activated protein kinase (AMPK) and insulin receptor/Akt signaling. here, we have studied the role of PIKE proteins in metabolic regulation in the fly. We show that the Drosophila PIKE homolog, ceng1A, encodes functional GTPases whose internal GAP domains catalyze their GTPase activity. To elucidate the biological function of ceng1A in flies, we introduced a deletion in the ceng1A gene by homologous recombination that removes all predicted functional PIKE domains. We found that homozygous ceng1A mutant animals survive to adulthood. In contrast to PIKE -/- mouse mutants, genetic ablation of Drosophila ceng1A does not result in growth defects or weight reduction. Although metabolic pathways such as insulin signaling, sensitivity towards starvation and mobilization of lipids under high fed conditions are not perturbed in ceng1A mutants, homozygous ceng1A mutants show a prolonged development in second instar larval stage, leading to a late onset of pupariation. In line with these results we found that expression of ecdysone inducible genes is reduced in ceng1A mutants. Together, we propose a novel role for Drosophila Ceng1A in regulating ecdysone signaling-dependent second to third instar larval transition.
Conflict of interest statement
Figures
Similar articles
-
Downregulation of Centaurin gamma1A increases synaptic transmission at Drosophila larval neuromuscular junctions.Eur J Neurosci. 2014 Oct;40(8):3158-70. doi: 10.1111/ejn.12681. Epub 2014 Jul 29. Eur J Neurosci. 2014. PMID: 25074496
-
Increased expression of the PI3K enhancer PIKE mediates deficits in synaptic plasticity and behavior in fragile X syndrome.Cell Rep. 2015 May 5;11(5):727-36. doi: 10.1016/j.celrep.2015.03.060. Epub 2015 Apr 23. Cell Rep. 2015. PMID: 25921541 Free PMC article.
-
rigor mortis encodes a novel nuclear receptor interacting protein required for ecdysone signaling during Drosophila larval development.Development. 2004 Jan;131(1):25-36. doi: 10.1242/dev.00920. Epub 2003 Nov 26. Development. 2004. PMID: 14645129
-
PIKE GTPase signaling and function.Int J Biol Sci. 2005;1(2):44-50. doi: 10.7150/ijbs.1.44. Epub 2005 Apr 1. Int J Biol Sci. 2005. PMID: 15951849 Free PMC article. Review.
-
PIKE GTPase: a novel mediator of phosphoinositide signaling.J Cell Sci. 2004 Jan 15;117(Pt 2):155-61. doi: 10.1242/jcs.00924. J Cell Sci. 2004. PMID: 14676271 Review.
Cited by
-
Ohgata, the Single Drosophila Ortholog of Human Cereblon, Regulates Insulin Signaling-dependent Organismic Growth.J Biol Chem. 2016 Nov 25;291(48):25120-25132. doi: 10.1074/jbc.M116.757823. Epub 2016 Oct 4. J Biol Chem. 2016. PMID: 27702999 Free PMC article.
-
Mitochondrial iron supply is required for the developmental pulse of ecdysone biosynthesis that initiates metamorphosis in Drosophila melanogaster.J Biol Inorg Chem. 2015 Dec;20(8):1229-38. doi: 10.1007/s00775-015-1302-2. Epub 2015 Oct 14. J Biol Inorg Chem. 2015. PMID: 26468126
-
The Genomic Basis of Postponed Senescence in Drosophila melanogaster.PLoS One. 2015 Sep 17;10(9):e0138569. doi: 10.1371/journal.pone.0138569. eCollection 2015. PLoS One. 2015. PMID: 26378456 Free PMC article.
-
AGAP1-associated endolysosomal trafficking abnormalities link gene-environment interactions in a neurodevelopmental disorder.bioRxiv [Preprint]. 2023 Jan 31:2023.01.31.526497. doi: 10.1101/2023.01.31.526497. bioRxiv. 2023. Update in: Dis Model Mech. 2023 Sep 1;16(9):dmm049838. doi: 10.1242/dmm.049838. PMID: 36778426 Free PMC article. Updated. Preprint.
-
AGAP1-associated endolysosomal trafficking abnormalities link gene-environment interactions in neurodevelopmental disorders.Dis Model Mech. 2023 Sep 1;16(9):dmm049838. doi: 10.1242/dmm.049838. Epub 2023 Sep 26. Dis Model Mech. 2023. PMID: 37470098 Free PMC article.
References
-
- Rong R, Ahn J, Huang H, Nagata E, Kalman D, et al. (2003) PI3 kinase enhancer-Homer complex couples mGluRI to PI3 kinase, preventing neuronal apoptosis. Nat. . Neurosci. 6 (11): 1153–1161. - PubMed
-
- Ahn J, Rong R, Kroll TG, van Meir EG, Snyder SH, et al. (2004) PIKE (phosphatidylinositol 3-kinase enhancer)-A GTPase stimulates Akt activity and mediates cellular invasion. . J. Biol. Chem. 279 (16): 16441–16451. - PubMed
-
- Reiser G, Bernstein H (2004) Altered expression of protein p42IP4/centaurin-alpha 1 in Alzheimer's disease brains and possible interaction of p42IP4 with nucleolin. Neuroreport 15 (1): 147–148. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

