Genome-wide analysis of cold adaptation in indigenous Siberian populations
- PMID: 24847810
- PMCID: PMC4029955
- DOI: 10.1371/journal.pone.0098076
Genome-wide analysis of cold adaptation in indigenous Siberian populations
Abstract
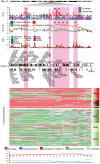
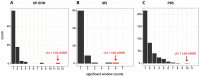
Following the dispersal out of Africa, where hominins evolved in warm environments for millions of years, our species has colonised different climate zones of the world, including high latitudes and cold environments. The extent to which human habitation in (sub-)Arctic regions has been enabled by cultural buffering, short-term acclimatization and genetic adaptations is not clearly understood. Present day indigenous populations of Siberia show a number of phenotypic features, such as increased basal metabolic rate, low serum lipid levels and increased blood pressure that have been attributed to adaptation to the extreme cold climate. In this study we introduce a dataset of 200 individuals from ten indigenous Siberian populations that were genotyped for 730,525 SNPs across the genome to identify genes and non-coding regions that have undergone unusually rapid allele frequency and long-range haplotype homozygosity change in the recent past. At least three distinct population clusters could be identified among the Siberians, each of which showed a number of unique signals of selection. A region on chromosome 11 (chr11:66-69 Mb) contained the largest amount of clustering of significant signals and also the strongest signals in all the different selection tests performed. We present a list of candidate cold adaption genes that showed significant signals of positive selection with our strongest signals associated with genes involved in energy regulation and metabolism (CPT1A, LRP5, THADA) and vascular smooth muscle contraction (PRKG1). By employing a new method that paints phased chromosome chunks by their ancestry we distinguish local Siberian-specific long-range haplotype signals from those introduced by admixture.
Conflict of interest statement
Figures

References
-
- Goebel T (1999) Pleistocene human colonization of Siberia and peopling of the Americas: An ecological approach. Evolutionary Anthropology: Issues, News, and Reviews 8: 208–227.
-
- Goebel T, Waters MR, O'Rourke DH (2008) The Late Pleistocene Dispersal of Modern Humans in the Americas. Science 319: 1497–1502. - PubMed
-
- Pitulko VV, Nikolsky PA, Girya EY, Basilyan AE, Tumskoy VE, et al. (2004) The Yana RHS site: humans in the Arctic before the last glacial maximum. Science 303: 52–56. - PubMed
-
- Clark PU, Dyke AS, Shakun JD, Carlson AE, Clark J, et al. (2009) The Last Glacial Maximum. Science 325: 710–714. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

