A comprehensive protein-protein interactome for yeast PAS kinase 1 reveals direct inhibition of respiration through the phosphorylation of Cbf1
- PMID: 24850888
- PMCID: PMC4091833
- DOI: 10.1091/mbc.E13-10-0631
A comprehensive protein-protein interactome for yeast PAS kinase 1 reveals direct inhibition of respiration through the phosphorylation of Cbf1
Abstract
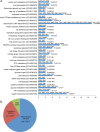
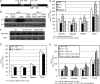
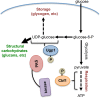
Per-Arnt-Sim (PAS) kinase is a sensory protein kinase required for glucose homeostasis in yeast, mice, and humans, yet little is known about the molecular mechanisms of its function. Using both yeast two-hybrid and copurification approaches, we identified the protein-protein interactome for yeast PAS kinase 1 (Psk1), revealing 93 novel putative protein binding partners. Several of the Psk1 binding partners expand the role of PAS kinase in glucose homeostasis, including new pathways involved in mitochondrial metabolism. In addition, the interactome suggests novel roles for PAS kinase in cell growth (gene/protein expression, replication/cell division, and protein modification and degradation), vacuole function, and stress tolerance. In vitro kinase studies using a subset of 25 of these binding partners identified Mot3, Zds1, Utr1, and Cbf1 as substrates. Further evidence is provided for the in vivo phosphorylation of Cbf1 at T211/T212 and for the subsequent inhibition of respiration. This respiratory role of PAS kinase is consistent with the reported hypermetabolism of PAS kinase-deficient mice, identifying a possible molecular mechanism and solidifying the evolutionary importance of PAS kinase in the regulation of glucose homeostasis.
© 2014 DeMille et al. This article is distributed by The American Society for Cell Biology under license from the author(s). Two months after publication it is available to the public under an Attribution–Noncommercial–Share Alike 3.0 Unported Creative Commons License (http://creativecommons.org/licenses/by-nc-sa/3.0).
Figures


Similar articles
-
The Regulation of Cbf1 by PAS Kinase Is a Pivotal Control Point for Lipogenesis vs. Respiration in Saccharomyces cerevisiae.G3 (Bethesda). 2019 Jan 9;9(1):33-46. doi: 10.1534/g3.118.200663. G3 (Bethesda). 2019. PMID: 30381292 Free PMC article.
-
PAS kinase is activated by direct SNF1-dependent phosphorylation and mediates inhibition of TORC1 through the phosphorylation and activation of Pbp1.Mol Biol Cell. 2015 Feb 1;26(3):569-82. doi: 10.1091/mbc.E14-06-1088. Epub 2014 Nov 26. Mol Biol Cell. 2015. PMID: 25428989 Free PMC article.
-
Regulation and function of yeast PAS kinase: a role in the maintenance of cellular integrity.Cell Cycle. 2009 Jun 15;8(12):1824-32. doi: 10.4161/cc.8.12.8799. Epub 2009 Jun 20. Cell Cycle. 2009. PMID: 19440050 Free PMC article.
-
PAS kinase: a nutrient sensing regulator of glucose homeostasis.IUBMB Life. 2013 Nov;65(11):921-9. doi: 10.1002/iub.1219. Epub 2013 Nov 7. IUBMB Life. 2013. PMID: 24265199 Free PMC article. Review.
-
New functions of protein kinase Gcn2 in yeast and mammals.IUBMB Life. 2012 Dec;64(12):971-4. doi: 10.1002/iub.1090. Epub 2012 Nov 5. IUBMB Life. 2012. PMID: 23129244 Review.
Cited by
-
Gut Microbiota Regulates the Interaction between Diet and Genetics to Influence Glucose Tolerance.Medicines (Basel). 2021 Jul 1;8(7):34. doi: 10.3390/medicines8070034. Medicines (Basel). 2021. PMID: 34357150 Free PMC article.
-
Next-generation gene discovery for variants of large impact on lipid traits.Curr Opin Lipidol. 2015 Apr;26(2):114-9. doi: 10.1097/MOL.0000000000000156. Curr Opin Lipidol. 2015. PMID: 25636063 Free PMC article. Review.
-
Per-Arnt-Sim Kinase (PASK): An Emerging Regulator of Mammalian Glucose and Lipid Metabolism.Nutrients. 2015 Sep 7;7(9):7437-50. doi: 10.3390/nu7095347. Nutrients. 2015. PMID: 26371032 Free PMC article. Review.
-
Ataxin-2: From RNA Control to Human Health and Disease.Genes (Basel). 2017 Jun 5;8(6):157. doi: 10.3390/genes8060157. Genes (Basel). 2017. PMID: 28587229 Free PMC article. Review.
-
Per-Arnt-Sim Kinase (PASK) Deficiency Increases Cellular Respiration on a Standard Diet and Decreases Liver Triglyceride Accumulation on a Western High-Fat High-Sugar Diet.Nutrients. 2018 Dec 15;10(12):1990. doi: 10.3390/nu10121990. Nutrients. 2018. PMID: 30558306 Free PMC article.
References
-
- Amberg DC, Burke D, Strathern JN. Methods in Yeast Genetics: A Cold Spring Harbor Laboratory Course Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 2005.
-
- An R, da Silva Xavier G, Hao HX, Semplici F, Rutter J, Rutter GA. Regulation by Per-Arnt-Sim (PAS) kinase of pancreatic duodenal homeobox-1 nuclear import in pancreatic beta-cells. Biochem Soc Trans. 2006;34:791–793. - PubMed
-
- Auer S, Hahne P, Soyal SM, Felder T, Miller K, Paulmichl M, Krempler F, Oberkofler H, Patsch W. Potential role of upstream stimulatory factor 1 gene variant in familial combined hyperlipidemia and related disorders. Arterioscler Thromb Vasc Biol. 2012;32:1535–1544. - PubMed
-
- Baker RE, Fitzgerald-Hayes M, O'Brien TC. Purification of the yeast centromere binding protein CP1 and a mutational analysis of its binding site. J Biol Chem. 1989;264:10843–10850. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

