eIF2B is a decameric guanine nucleotide exchange factor with a γ2ε2 tetrameric core
- PMID: 24852487
- PMCID: PMC4046112
- DOI: 10.1038/ncomms4902
eIF2B is a decameric guanine nucleotide exchange factor with a γ2ε2 tetrameric core
Abstract
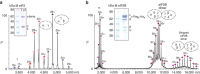
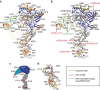
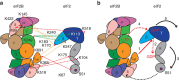
eIF2B facilitates and controls protein synthesis in eukaryotes by mediating guanine nucleotide exchange on its partner eIF2. We combined mass spectrometry (MS) with chemical cross-linking, surface accessibility measurements and homology modelling to define subunit stoichiometry and interactions within eIF2B and eIF2. Although it is generally accepted that eIF2B is a pentamer of five non-identical subunits (α-ε), here we show that eIF2B is a decamer. MS and cross-linking of eIF2B complexes allows us to propose a model for the subunit arrangements within eIF2B where the subunit assembly occurs through catalytic γ- and ε-subunits, with regulatory subunits arranged in asymmetric trimers associated with the core. Cross-links between eIF2 and eIF2B allow modelling of interactions that contribute to nucleotide exchange and its control by eIF2 phosphorylation. Finally, we identify that GTP binds to eIF2Bγ, prompting us to propose a multi-step mechanism for nucleotide exchange.
Figures



Similar articles
-
The beta/Gcd7 subunit of eukaryotic translation initiation factor 2B (eIF2B), a guanine nucleotide exchange factor, is crucial for binding eIF2 in vivo.Mol Cell Biol. 2010 Nov;30(21):5218-33. doi: 10.1128/MCB.00265-10. Epub 2010 Aug 30. Mol Cell Biol. 2010. PMID: 20805354 Free PMC article.
-
Identification of domains and residues within the epsilon subunit of eukaryotic translation initiation factor 2B (eIF2Bepsilon) required for guanine nucleotide exchange reveals a novel activation function promoted by eIF2B complex formation.Mol Cell Biol. 2000 Jun;20(11):3965-76. doi: 10.1128/MCB.20.11.3965-3976.2000. Mol Cell Biol. 2000. PMID: 10805739 Free PMC article.
-
Crystal structure of eukaryotic translation initiation factor 2B.Nature. 2016 Mar 3;531(7592):122-5. doi: 10.1038/nature16991. Epub 2016 Feb 22. Nature. 2016. PMID: 26901872
-
Crystal structure of eIF2B and insights into eIF2-eIF2B interactions.FEBS J. 2017 Mar;284(6):868-874. doi: 10.1111/febs.13896. Epub 2016 Sep 29. FEBS J. 2017. PMID: 27627185 Review.
-
A new function and complexity for protein translation initiation factor eIF2B.Cell Cycle. 2014;13(17):2660-5. doi: 10.4161/15384101.2014.948797. Cell Cycle. 2014. PMID: 25486352 Free PMC article. Review.
Cited by
-
Pharmacological dimerization and activation of the exchange factor eIF2B antagonizes the integrated stress response.Elife. 2015 Apr 15;4:e07314. doi: 10.7554/eLife.07314. Elife. 2015. PMID: 25875391 Free PMC article.
-
Surviving and Adapting to Stress: Translational Control and the Integrated Stress Response.Antioxid Redox Signal. 2023 Aug;39(4-6):351-373. doi: 10.1089/ars.2022.0123. Epub 2023 May 9. Antioxid Redox Signal. 2023. PMID: 36943285 Free PMC article. Review.
-
Vanishing white matter: Eukaryotic initiation factor 2B model and the impact of missense mutations.Mol Genet Genomic Med. 2021 Mar;9(3):e1593. doi: 10.1002/mgg3.1593. Epub 2021 Jan 12. Mol Genet Genomic Med. 2021. PMID: 33432707 Free PMC article.
-
From molecular chaperones to membrane motors: through the lens of a mass spectrometrist.Biochem Soc Trans. 2017 Feb 8;45(1):251-260. doi: 10.1042/BST20160395. Biochem Soc Trans. 2017. PMID: 28202679 Free PMC article.
-
Blind testing cross-linking/mass spectrometry under the auspices of the 11th critical assessment of methods of protein structure prediction (CASP11).Wellcome Open Res. 2016 Dec 9;1:24. doi: 10.12688/wellcomeopenres.10046.1. Wellcome Open Res. 2016. PMID: 28317030 Free PMC article.
References
-
- Maag D., Fekete C. A., Gryczynski Z. & Lorsch J. R. A conformational change in the eukaryotic translation preinitiation complex and release of eIF1 signal recognition of the start codon. Mol. Cell 17, 265–275 (2005). - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- MC_U105184332/MRC_/Medical Research Council/United Kingdom
- BB/F013272/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- BB/H010599/1/BB_/Biotechnology and Biological Sciences Research Council/United Kingdom
- 088150/B/09/Z/WT_/Wellcome Trust/United Kingdom
- 088150/WT_/Wellcome Trust/United Kingdom
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

