Conformational switching in PolyGln amyloid fibrils resulting from a single amino acid insertion
- PMID: 24853742
- PMCID: PMC4052364
- DOI: 10.1016/j.bpj.2014.03.047
Conformational switching in PolyGln amyloid fibrils resulting from a single amino acid insertion
Abstract
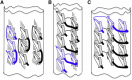
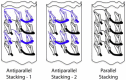
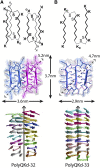
The established correlation between neurodegenerative disorders and intracerebral deposition of polyglutamine aggregates motivates attempts to better understand their fibrillar structure. We designed polyglutamines with a few lysines inserted to overcome the hindrance of extreme insolubility and two D-lysines to limit the lengths of β-strands. One is 33 amino acids long (PolyQKd-33) and the other has one fewer glutamine (PolyQKd-32). Both form well-dispersed fibrils suitable for analysis by electron microscopy. Electron diffraction confirmed cross-β structures in both fibrils. Remarkably, the deletion of just one glutamine residue from the middle of the peptide leads to substantially different amyloid structures. PolyQKd-32 fibrils are consistently 10-20% wider than PolyQKd-33, as measured by negative staining, cryo-electron microscopy, and scanning transmission electron microscopy. Scanning transmission electron microscopy analysis revealed that the PolyQKd-32 fibrils have 50% higher mass-per-length than PolyQKd-33. This distinction can be explained by a superpleated β-structure model for PolyQKd-33 and a model with two β-solenoid protofibrils for PolyQKd-32. These data provide evidence for β-arch-containing structures in polyglutamine fibrils and open future possibilities for structure-based drug design.
Copyright © 2014 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures





Similar articles
-
Amyloid-like features of polyglutamine aggregates and their assembly kinetics.Biochemistry. 2002 Jun 11;41(23):7391-9. doi: 10.1021/bi011772q. Biochemistry. 2002. PMID: 12044172
-
α-Synuclein Amyloid Fibrils with Two Entwined, Asymmetrically Associated Protofibrils.J Biol Chem. 2016 Jan 29;291(5):2310-8. doi: 10.1074/jbc.M115.698787. Epub 2015 Dec 7. J Biol Chem. 2016. PMID: 26644467 Free PMC article.
-
Mapping abeta amyloid fibril secondary structure using scanning proline mutagenesis.J Mol Biol. 2004 Jan 16;335(3):833-42. doi: 10.1016/j.jmb.2003.11.008. J Mol Biol. 2004. PMID: 14687578
-
Molecular structures of amyloid and prion fibrils: consensus versus controversy.Acc Chem Res. 2013 Jul 16;46(7):1487-96. doi: 10.1021/ar300282r. Epub 2013 Jan 7. Acc Chem Res. 2013. PMID: 23294335 Free PMC article. Review.
-
Alzheimer's amyloid fibrils: structure and assembly.Biochim Biophys Acta. 2000 Jul 26;1502(1):16-30. doi: 10.1016/s0925-4439(00)00029-6. Biochim Biophys Acta. 2000. PMID: 10899428 Review.
Cited by
-
W8, a new Sup35 prion strain, transmits distinctive information with a conserved assembly scheme.Prion. 2015;9(3):207-27. doi: 10.1080/19336896.2015.1039217. Prion. 2015. PMID: 26038983 Free PMC article.
-
Proteomic Evidence for Amyloidogenic Cross-Seeding in Fibrinaloid Microclots.Int J Mol Sci. 2024 Oct 8;25(19):10809. doi: 10.3390/ijms251910809. Int J Mol Sci. 2024. PMID: 39409138 Free PMC article.
References
-
- Jorda J., Kajava A.V. Protein homorepeats sequences, structures, evolution, and functions. Adv. Protein Chem. Struct. Biol. 2010;79:59–88. - PubMed
-
- Orr H.T., Zoghbi H.Y. Trinucleotide repeat disorders. Annu. Rev. Neurosci. 2007;30:575–621. - PubMed
-
- Davies S.W., Turmaine M., Bates G.P. Formation of neuronal intranuclear inclusions underlies the neurological dysfunction in mice transgenic for the HD mutation. Cell. 1997;90:537–548. - PubMed
-
- Becher M.W., Kotzuk J.A., Ross C.A. Intranuclear neuronal inclusions in Huntington’s disease and dentatorubral and pallidoluysian atrophy: correlation between the density of inclusions and IT15 CAG triplet repeat length. Neurobiol. Dis. 1998;4:387–397. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

