Mouse knockout models for HIV-1 restriction factors
- PMID: 24854580
- PMCID: PMC4160573
- DOI: 10.1007/s00018-014-1646-8
Mouse knockout models for HIV-1 restriction factors
Abstract
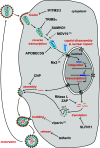
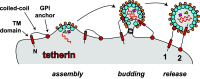
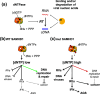
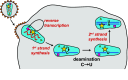
Infection of cells with human immunodeficiency virus 1 (HIV-1) is controlled by restriction factors, host proteins that counteract a variety of steps in the life cycle of this lentivirus. These include SAMHD1, APOBEC3G and tetherin, which block reverse transcription, hypermutate viral DNA and prevent progeny virus release, respectively. These and other HIV-1 restriction factors are conserved and have clear orthologues in the mouse. This review summarises studies in knockout mice lacking HIV-1 restriction factors. In vivo experiments in such animals have not only validated in vitro data obtained from cultured cells, but have also revealed new findings about the biology of these proteins. Indeed, genetic ablation of HIV-1 restriction factors in the mouse has provided evidence that restriction factors control retroviruses and other viruses in vivo and has led to new insights into the mechanisms by which these proteins counteract infection. For example, in vivo experiments in knockout mice demonstrate that virus control exerted by restriction factors can shape adaptive immune responses. Moreover, the availability of animals lacking restriction factors opens the possibility to study the function of these proteins in other contexts such as autoimmunity and cancer. Further in vivo studies of more recently identified HIV-1 restriction factors in gene targeted mice are, therefore, justified.
Figures
Similar articles
-
The restriction factors of human immunodeficiency virus.J Biol Chem. 2012 Nov 30;287(49):40875-83. doi: 10.1074/jbc.R112.416925. Epub 2012 Oct 5. J Biol Chem. 2012. PMID: 23043100 Free PMC article. Review.
-
SAMHD1-dependent retroviral control and escape in mice.EMBO J. 2013 Sep 11;32(18):2454-62. doi: 10.1038/emboj.2013.163. Epub 2013 Jul 19. EMBO J. 2013. PMID: 23872947 Free PMC article.
-
SAMHD1 knockout mice: modeling retrovirus restriction in vivo.Retrovirology. 2013 Nov 20;10:142. doi: 10.1186/1742-4690-10-142. Retrovirology. 2013. PMID: 24257155 Free PMC article.
-
Intrinsic host restrictions to HIV-1 and mechanisms of viral escape.Nat Immunol. 2015 Jun;16(6):546-53. doi: 10.1038/ni.3156. Nat Immunol. 2015. PMID: 25988886 Free PMC article. Review.
-
Intrinsic cellular defenses against human immunodeficiency viruses.Immunity. 2012 Sep 21;37(3):399-411. doi: 10.1016/j.immuni.2012.08.013. Immunity. 2012. PMID: 22999946 Free PMC article.
Cited by
-
HEK293T cell lines defective for O-linked glycosylation.PLoS One. 2017 Jun 27;12(6):e0179949. doi: 10.1371/journal.pone.0179949. eCollection 2017. PLoS One. 2017. PMID: 28654657 Free PMC article.
-
Reconstruction of a replication-competent ancestral murine endogenous retrovirus-L.Retrovirology. 2018 May 2;15(1):34. doi: 10.1186/s12977-018-0416-3. Retrovirology. 2018. PMID: 29716624 Free PMC article.
-
Nucleic acid recognition orchestrates the anti-viral response to retroviruses.Cell Host Microbe. 2015 Apr 8;17(4):478-88. doi: 10.1016/j.chom.2015.02.021. Epub 2015 Mar 26. Cell Host Microbe. 2015. PMID: 25816774 Free PMC article.
-
MicroRNA-mediated regulation of p21 and TASK1 cellular restriction factors enhances HIV-1 infection.J Cell Sci. 2015 Apr 15;128(8):1607-16. doi: 10.1242/jcs.167817. Epub 2015 Feb 25. J Cell Sci. 2015. PMID: 25717002 Free PMC article.
-
The C-terminal domain of feline and bovine SAMHD1 proteins has a crucial role in lentiviral restriction.J Biol Chem. 2020 Mar 27;295(13):4252-4264. doi: 10.1074/jbc.RA120.012767. Epub 2020 Feb 19. J Biol Chem. 2020. PMID: 32075911 Free PMC article.
References
-
- Sorek R, Lawrence CM, Wiedenheft B. CRISPR-mediated adaptive immune systems in bacteria and archaea. Annu Rev Biochem. 2013;82:237–266. - PubMed
-
- Nayak A, Tassetto M, Kunitomi M, Andino R. RNA interference-mediated intrinsic antiviral immunity in invertebrates. Curr Top Microbiol Immunol. 2013;371:183–200. - PubMed
-
- Szittya G, Burgyán J. RNA interference-mediated intrinsic antiviral immunity in plants. Curr Top Microbiol Immunol. 2013;371:153–181. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

