An essential mesenchymal function for miR-143/145 in intestinal epithelial regeneration
- PMID: 24855947
- PMCID: PMC4175516
- DOI: 10.1016/j.cell.2014.03.055
An essential mesenchymal function for miR-143/145 in intestinal epithelial regeneration
Abstract
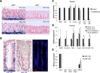
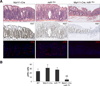
Downregulation of the miR-143/145 microRNA (miRNA) cluster has been repeatedly reported in colon cancer and other epithelial tumors. In addition, overexpression of these miRNAs inhibits tumorigenesis, leading to broad consensus that they function as cell-autonomous epithelial tumor suppressors. We generated mice with deletion of miR-143/145 to investigate the functions of these miRNAs in intestinal physiology and disease in vivo. Although intestinal development proceeded normally in the absence of these miRNAs, epithelial regeneration after injury was dramatically impaired. Surprisingly, we found that miR-143/145 are expressed and function exclusively within the mesenchymal compartment of intestine. Defective epithelial regeneration in miR-143/145-deficient mice resulted from the dysfunction of smooth muscle and myofibroblasts and was associated with derepression of the miR-143 target Igfbp5, which impaired IGF signaling after epithelial injury. These results provide important insights into the regulation of epithelial wound healing and argue against a cell-autonomous tumor suppressor role for miR-143/145 in colon cancer.
Copyright © 2014 Elsevier Inc. All rights reserved.
Figures





Comment in
-
Seeing through the miRage of tissue complexity.Cell Cycle. 2014;13(19):2988-9. doi: 10.4161/15384101.2014.962852. Cell Cycle. 2014. PMID: 25486556 Free PMC article. No abstract available.
References
-
- Andoh A, Bamba S, Brittan M, Fujiyama Y, Wright NA. Role of intestinal subepithelial myofibroblasts in inflammation and regenerative response in the gut. Pharmacology & Therapeutics. 2007;114:94–106. - PubMed
-
- Barker N, van Es JH, Kuipers J, Kujala P, van den Born M, Cozijnsen M, Haegebarth A, Korving J, Begthel H, Peters PJ, et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature. 2007;449:1003–1007. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

