Characterization of the canine mda-7 gene, transcripts and expression patterns
- PMID: 24865935
- PMCID: PMC4131717
- DOI: 10.1016/j.gene.2014.05.054
Characterization of the canine mda-7 gene, transcripts and expression patterns
Abstract
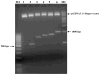
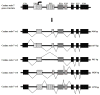
Human melanoma differentiation associated gene-7/interleukin-24 (mda-7/IL-24) displays potent growth suppressing and cell killing activity against a wide variety of human and rodent cancer cells. In this study, we identified a canine ortholog of the human mda-7/IL-24 gene located within a cluster of IL-10 family members on chromosome 7. The full-length mRNA sequence of canine mda-7 was determined, which encodes a 186-amino acid protein that has 66% similarity to human MDA-7/IL-24. Canine MDA-7 is constitutively expressed in cultured normal canine epidermal keratinocytes (NCEKs), and its expression levels are increased after lipopolysaccharide stimulation. In cultured NCEKs, the canine mda-7 pre-mRNA is differentially spliced, via exon skipping and alternate 5'-splice donor sites, to yield five splice variants (canine mda-7sv1, canine mda-7sv2, canine mda-7sv3, canine mda-7sv4 and canine mda-7sv5) that encode four protein isoforms of the canine MDA-7 protein. These protein isoforms have a conserved N-terminus (signal peptide sequence) and are dissimilar in amino acid sequences at their C-terminus. Canine MDA-7 is not expressed in primary canine tumor samples, and most tumor derived cancer cell lines tested, like its human counterpart. Unlike human MDA-7/IL-24, canine mda-7 mRNA is not expressed in unstimulated or lipopolysaccharide (LPS), concanavalin A (ConA) or phytohemagglutinin (PHA) stimulated canine peripheral blood mononuclear cells (PBMCs). Furthermore, in-silico analysis revealed that canonical canine MDA-7 has a potential 28 amino acid signal peptide sequence that can target it for active secretion. This data suggests that canine mda-7 is indeed an ortholog of human mda-7/IL-24, its protein product has high amino acid similarity to human MDA-7/IL-24 protein and it may possess similar biological properties to human MDA-7/IL-24, but its expression pattern is more restricted than its human ortholog.
Keywords: Interleukin-24 (IL-24); Melanoma differentiation associated gene-7 (mda-7); cancer cell lines; canine.
Copyright © 2014 Elsevier B.V. All rights reserved.
Figures

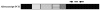
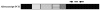


Similar articles
-
Identification of interleukin-26 in the dromedary camel (Camelus dromedarius): Evidence of alternative splicing and isolation of novel splice variants.Mol Immunol. 2015 Oct;67(2 Pt B):357-68. doi: 10.1016/j.molimm.2015.06.022. Epub 2015 Jul 17. Mol Immunol. 2015. PMID: 26190308 Free PMC article.
-
Alternative splicing of IL-24 in melanocytes by deletion of exons 3 and 5.Int J Immunogenet. 2005 Dec;32(6):375-8. doi: 10.1111/j.1744-313X.2005.00540.x. Int J Immunogenet. 2005. PMID: 16313301
-
Loss of novel mda-7 splice variant (mda-7s) expression is associated with metastatic melanoma.J Invest Dermatol. 2004 Sep;123(3):583-8. doi: 10.1111/j.0022-202X.2004.23321.x. J Invest Dermatol. 2004. PMID: 15304100
-
MDA-7/IL-24: novel cancer growth suppressing and apoptosis inducing cytokine.Cytokine Growth Factor Rev. 2003 Feb;14(1):35-51. doi: 10.1016/s1359-6101(02)00074-6. Cytokine Growth Factor Rev. 2003. PMID: 12485618 Review.
-
MDA-7/IL-24 is a unique cytokine--tumor suppressor in the IL-10 family.Int Immunopharmacol. 2004 May;4(5):649-67. doi: 10.1016/j.intimp.2004.01.017. Int Immunopharmacol. 2004. PMID: 15120650 Review.
Cited by
-
Insights into the Mechanisms of Action of MDA-7/IL-24: A Ubiquitous Cancer-Suppressing Protein.Int J Mol Sci. 2021 Dec 22;23(1):72. doi: 10.3390/ijms23010072. Int J Mol Sci. 2021. PMID: 35008495 Free PMC article. Review.
References
-
- Allen M, Pratscher B, Krepler C, Frei K, Schofer C, Pehamberger H, Muller M, Lucas T. Alternative splicing of IL-24 in melanocytes by deletion of exons 3 and 5. Int J Immunogenet. 2005;32:375–8. - PubMed
-
- Allen M, Pratscher B, Roka F, Krepler C, Wacheck V, Schofer C, Pehamberger H, Muller M, Lucas T. Loss of novel mda-7 splice variant (mda-7s) expression is associated with metastatic melanoma. J Invest Dermatol. 2004;123:583–8. - PubMed
-
- Caudell EG, Mumm JB, Poindexter N, Ekmekcioglu S, Mhashilkar AM, Yang XH, Retter MW, Hill P, Chada S, Grimm EA. The protein product of the tumor suppressor gene, melanoma differentiation-associated gene 7, exhibits immunostimulatory activity and is designated IL-24. J Immunol. 2002;168:6041–6. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

