Substrate specificity of the lanthipeptide peptidase ElxP and the oxidoreductase ElxO
- PMID: 24866416
- PMCID: PMC4136673
- DOI: 10.1021/cb5002526
Substrate specificity of the lanthipeptide peptidase ElxP and the oxidoreductase ElxO
Abstract
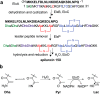
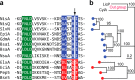
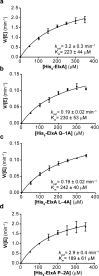
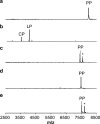
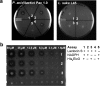
The final step in lanthipeptide biosynthesis involves the proteolytic removal of an N-terminal leader peptide. In the class I lanthipeptide epilancin 15X, this step is performed by the subtilisin-like serine peptidase ElxP. Bioinformatic, kinetic, and mass spectrometric analysis revealed that ElxP recognizes the stretch of amino acids DLNPQS located near the proteolytic cleavage site of its substrate, ElxA. When the ElxP recognition motif was inserted into the noncognate lanthipeptide precursor NisA, ElxP was able to proteolytically remove the leader peptide from NisA. Proteolytic removal of the leader peptide by ElxP during the biosynthesis of epilancin 15X exposes an N-terminal dehydroalanine on the core peptide of ElxA that hydrolyzes to a pyruvyl group. The short-chain dehydrogenase ElxO reduces the pyruvyl group to a lactyl moiety in the final step of epilancin 15X maturation. Using synthetic peptides, we also investigated the substrate specificity of ElxO and determined the 1.85 Å resolution X-ray crystal structure of the enzyme.
Figures

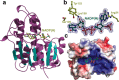
References
-
- Arnison P. G.; Bibb M. J.; Bierbaum G.; Bowers A. A.; Bugni T. S.; Bulaj G.; Camarero J. A.; Campopiano D. J.; Challis G. L.; Clardy J.; Cotter P. D.; Craik D. J.; Dawson M.; Dittmann E.; Donadio S.; Dorrestein P. C.; Entian K.-D.; Fischbach M. A.; Garavelli J. S.; Göransson U.; Gruber C. W.; Haft D. H.; Hemscheidt T. K.; Hertweck C.; Hill C.; Horswill A. R.; Jaspars M.; Kelly W. L.; Klinman J. P.; Kuipers O. P.; Link A. J.; Liu W.; Marahiel M. A.; Mitchell D. A.; Moll G. N.; Moore B. S.; Müller R.; Nair S. K.; Nes I. F.; Norris G. E.; Olivera B. M.; Onaka H.; Patchett M. L.; Piel J.; Reaney M. J. T.; Rebuffat S.; Ross R. P.; Sahl H.-G.; Schmidt E. W.; Selsted M. E.; Severinov K.; Shen B.; Sivonen K.; Smith L.; Stein T.; Süssmuth R. E.; Tagg J. R.; Tang G.-L.; Truman A. W.; Vederas J. C.; Walsh C. T.; Walton J. D.; Wenzel S. C.; Willey J. M.; van der Donk W. A. (2013) Ribosomally synthesized and post-translationally modified peptide natural products: Overview and recommendations for a universal nomenclature. Nat. Prod. Rep. 30, 108–160. - PMC - PubMed
-
- Schnell N.; Entian K.-D.; Schneider U.; Götz F.; Zahner H.; Kellner R.; Jung G. (1988) Prepeptide sequence of epidermin, a ribosomally synthesized antibiotic with four sulphide-rings. Nature 333, 276–278. - PubMed
-
- van der Meer J. R.; Rollema H. S.; Siezen R. J.; Beerthuyzen M. M.; Kuipers O. P.; de Vos W. M. (1994) Influence of amino acid substitutions in the nisin leader peptide on biosynthesis and secretion of nisin by Lactococcus lactis. J. Biol. Chem. 269, 3555–3562. - PubMed
-
- Li B.; Yu J. P.; Brunzelle J. S.; Moll G. N.; van der Donk W. A.; Nair S. K. (2006) Structure and mechanism of the lantibiotic cyclase involved in nisin biosynthesis. Science 311, 1464–1467. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

