Gene expression profiling reveals large regulatory switches between succeeding stipe stages in Volvariella volvacea
- PMID: 24867220
- PMCID: PMC4035324
- DOI: 10.1371/journal.pone.0097789
Gene expression profiling reveals large regulatory switches between succeeding stipe stages in Volvariella volvacea
Abstract
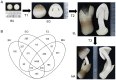
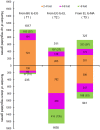
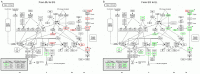
The edible mushroom Volvariella volvacea is an important crop in Southeast Asia and is predominantly harvested in the egg stage. One of the main factors that negatively affect its yield and value is the rapid transition from the egg to the elongation stage, which has a decreased commodity value and shelf life. To improve our understanding of the changes during stipe development and the transition from egg to elongation stage in particular, we analyzed gene transcription in stipe tissue of V. volvacea using 3'-tag based digital expression profiling. Stipe development turned out to be fairly complex with high numbers of expressed genes, and regulation of stage differences is mediated mainly by changes in expression levels of genes, rather than on/off modulation. Most explicit is the strong up-regulation of cell division from button to egg, and the very strong down-regulation hereof from egg to elongation, that continues in the maturation stage. Button and egg share cell division as means of growth, followed by a major developmental shift towards rapid stipe elongation based on cell extension as demonstrated by inactivation of cell division throughout elongation and maturation. Examination of regulatory genes up-regulated from egg to elongation identified three potential high upstream regulators for this switch. The new insights in stipe dynamics, together with a series of new target genes, will provide a sound base for further studies on the developmental mechanisms of mushroom stipes and the switch from egg to elongation in V. volvacea in particular.
Conflict of interest statement
Figures

Similar articles
-
Identification and expression analysis of a new glycoside hydrolase family 55 exo-β-1,3-glucanase-encoding gene in Volvariella volvacea suggests a role in fruiting body development.Gene. 2013 Sep 15;527(1):154-60. doi: 10.1016/j.gene.2013.05.071. Epub 2013 Jun 7. Gene. 2013. PMID: 23751305
-
Cloning and Expression Analysis of Vvlcc3, a Novel and Functional Laccase Gene Possibly Involved in Stipe Elongation.Int J Mol Sci. 2015 Dec 1;16(12):28498-509. doi: 10.3390/ijms161226111. Int J Mol Sci. 2015. PMID: 26633374 Free PMC article.
-
Characterization and expression pattern of homeobox transcription factors in fruiting body development of straw mushroom Volvariella volvacea.Fungal Biol. 2019 Feb;123(2):95-102. doi: 10.1016/j.funbio.2018.10.008. Epub 2018 Nov 12. Fungal Biol. 2019. PMID: 30709523
-
Analysis of synonymous codon usage patterns in the edible fungus Volvariella volvacea.Biotechnol Appl Biochem. 2017 Mar;64(2):218-224. doi: 10.1002/bab.1538. Epub 2016 Dec 15. Biotechnol Appl Biochem. 2017. PMID: 27696508 Review.
-
Increase in the incidence of invasive fungal infections due to Volvariella Volvacea.Eur J Clin Microbiol Infect Dis. 2024 May;43(5):1031-1036. doi: 10.1007/s10096-024-04800-3. Epub 2024 Mar 12. Eur J Clin Microbiol Infect Dis. 2024. PMID: 38472521 Review.
Cited by
-
Reactive Oxygen Species Distribution Involved in Stipe Gradient Elongation in the Mushroom Flammulina filiformis.Cells. 2022 Jun 11;11(12):1896. doi: 10.3390/cells11121896. Cells. 2022. PMID: 35741023 Free PMC article.
-
Transcriptome Profiling Reveals Candidate Genes Related to Stipe Gradient Elongation of Flammulina filiformis.J Fungi (Basel). 2022 Dec 31;9(1):64. doi: 10.3390/jof9010064. J Fungi (Basel). 2022. PMID: 36675885 Free PMC article.
-
Transcriptome data reveal conserved patterns of fruiting body development and response to heat stress in the mushroom-forming fungus Flammulina filiformis.PLoS One. 2020 Oct 16;15(10):e0239890. doi: 10.1371/journal.pone.0239890. eCollection 2020. PLoS One. 2020. PMID: 33064719 Free PMC article.
-
The Chromatin Modifier Protein FfJMHY Plays an Important Role in Regulating the Rate of Mycelial Growth and Stipe Elongation in Flammulina filiformis.J Fungi (Basel). 2022 May 3;8(5):477. doi: 10.3390/jof8050477. J Fungi (Basel). 2022. PMID: 35628733 Free PMC article.
-
Comparative Proteomic Analysis of Pleurotus ostreatus Reveals Great Metabolic Differences in the Cap and Stipe Development and the Potential Role of Ca2+ in the Primordium Differentiation.Int J Mol Sci. 2019 Dec 14;20(24):6317. doi: 10.3390/ijms20246317. Int J Mol Sci. 2019. PMID: 31847351 Free PMC article.
References
-
- Chang S (1977) The origin and early development of straw mushroom cultivation. Econ Bot 31: 374–376.
-
- Ding S, Ge W, Buswell JA (2007) Molecular cloning and transcriptional expression analysis of an intracellular β-glucosidase, a family 3 glycosyl hydrolase, from the edible straw mushroom, Volvariella volvacea . FEMS Microbiol Lett 267: 221–229. - PubMed
-
- Wang J, Guo L, Zhang K, Wu Q, Lin J (2008) Highly efficient Agrobacterium-mediated transformation of Volvariella volvacea . Bioresoure Technol 99: 8524–8527. - PubMed
-
- Mau JL, Chyau CC, Li JY, Tseng YH (1997) Flavor compounds in straw mushrooms Volvariella volvacea harvested at different stages of maturity. J Agr Food Chem 45: 4726–4729.
-
- Chang ST, Yau CK (1971) Volvariella volvacea and its life history. Am J Bot 58: 552–561.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

