RARRES3 suppresses breast cancer lung metastasis by regulating adhesion and differentiation
- PMID: 24867881
- PMCID: PMC4119352
- DOI: 10.15252/emmm.201303675
RARRES3 suppresses breast cancer lung metastasis by regulating adhesion and differentiation
Abstract
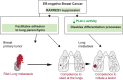
In estrogen receptor-negative breast cancer patients, metastatic relapse usually occurs in the lung and is responsible for the fatal outcome of the disease. Thus, a better understanding of the biology of metastasis is needed. In particular, biomarkers to identify patients that are at risk of lung metastasis could open the avenue for new therapeutic opportunities. Here we characterize the biological activity of RARRES3, a new metastasis suppressor gene whose reduced expression in the primary breast tumors identifies a subgroup of patients more likely to develop lung metastasis. We show that RARRES3 downregulation engages metastasis-initiating capabilities by facilitating adhesion of the tumor cells to the lung parenchyma. In addition, impaired tumor cell differentiation due to the loss of RARRES3 phospholipase A1/A2 activity also contributes to lung metastasis. Our results establish RARRES3 downregulation as a potential biomarker to identify patients at high risk of lung metastasis who might benefit from a differentiation treatment in the adjuvant programme.
Keywords: breast cancer; lung metastasis; metastasis suppressor.
© 2014 The Authors. Published under the terms of the CC BY 4.0 license.
Figures
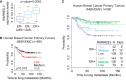
Box plot of RARRES3 expression levels in the MSKCC (n = 82) breast cancer tumor dataset according to ER and lung metastasis signature (LMS) status.
Kaplan–Meier representation of the probability of lung metastasis-free survival in the MSK/EMC breast cancer tumor dataset (n = 560) according to RARRES3 levels of expression. Low, medium, and high represent RARRES3 expression levels in the following way: low (< mean ± SD), medium (≥ mean ± SD and ≤ mean ± SD), and high (> mean ± SD).
Kaplan–Meier representation of the probability of lung metastasis-free survival in 560 breast cancer cases according to the ER status and RARRES 3 expression levels according to (B).

Schematic representation of in vivo experimental procedure to evaluate lung metastatic potential from the orthotopic site. The indicated cell lines (5 × 105 cells) were injected contralaterally into the fourth mammary fat pad of mice. Tumors reaching 300 mm3 were surgically removed. Seven days post-mastectomy, lung metastasis burden originated from size-matched tumors was quantified.
(Left panel) Quantification of ex vivo bioluminescent signal at the lungs in each experimental group subjected to the tumor growth/resection scheme described in (A) at end point. n = 8 and n = 9 mice per group were used. Whiskers plots from min–max values were used. (Right panel) Representative bioluminescence images of in vivo and ex vivo lung colonization of the mice are shown.
Representative human Vimentin IHC staining of whole lung sections to highlight metastatic tumor lesions from (B) are shown. Inset panels (4× magnification) reflect the size and multiple metastatic foci detected in the LM2-Mock group.
LM2 cells (5 × 105) expressing an empty vector (LM2-Mock) or a RARRES3-expressing vector (LM2-RARRES3) were injected contralaterally into the fourth mammary fat pad of mice, and tumor growth was measured over time. n = 20 per group. Data are averages ± SEM.

LM2-Mock and LM2-RARRES3 cells (2 × 105) were injected into the tail vein of mice. Lung colonization was assayed by weekly bioluminescence imaging. Plots show normalized photon flux in the lung over time (n = 10 per group). Representative H&E and Ki-67 staining of lung sections 4 weeks after engrafting are shown. Data are averages ± SEM.
Parental MDA-MB-231 cells (2 × 105) transduced with a control vector (shControl) or two independent RARRES3 shRNA vectors (shRARRES3 #1 and #2) were injected into the tail vein of mice. Lung colonization was assayed by weekly bioluminescence imaging. Plots show normalized photon flux in the lung over time (n = 10 per group). Representative H&E and Ki-67 staining of lung sections 4 weeks after xenografting are shown. Data are averages ± SEM.
CN37 patient-derived metastatic breast cancer cells (2 × 105) transduced with a control vector (shControl) or two independent RARRES3 shRNA vectors (shRARRES3 #1 and #2) were injected into the tail vein of mice (n = 8, 9 and 9 per group, respectively). Lung colonization was assayed by weekly bioluminescence imaging. Kaplan–Meier curve of the probability of lung metastasis-free survival for CN37 shControl, shRARRES3#1, and #2 is presented. Log-rank test was used to establish statistical significance.
4T1 mouse-derived breast cancer cells (2 × 105) transduced with a Mock or RARRES3 vector were injected into the tail vein of mice (n = 10 per group). Lung colonization was assayed by calculating the total area of lung metastasis lesion normalized per the total area of the lungs (H&E). Three sections were analyzed per lung. Data are averages of 10 lungs (mice) per group ± SEM. Wilcoxon test was used to establish statistical significance. Representative H&E sections are shown.
Vimentin IHC of lung sections from animals inoculated with the indicated lines. Upper panels: 2× magnification. Lower panels: 4× magnification. Quantification of the number of foci per field and of the average area per foci is shown in the right panels. Data are averages ± SEM.
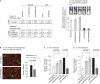
A (Left panel) Percentage of apoptotic cells under anoikis conditions, as determined by FACS measurement of Anexin V staining. Same number of cells of the indicated cell lines were plated, and measurements were taken at the indicated time point. Data are averages of three independent experiments ± SD. (Right panel) Apoptotic human luciferase activity of the indicated cell lines was measured in vivo using a Caspase-3 luciferin pro-substrate and normalized to total luciferase activity at the indicated time points after injection (n = 5 per group). Data are averages ± SEM. PI stands for post-injection.
B Cell tracker green-labeled LM2-mock and LM2-RARRES3 cell lines were injected via the tail vein and allowed to lodge in the lungs. Two day post-injection, mice were inoculated with rhodamine-lectin and 30 min later were perfused with 5 ml of PBS to remove cells attached to the vasculature. Lungs were extracted, flushed with PBS, and fixed-frozen in OCT, and frozen sections were obtained. Representative confocal images of extravasated cells (green) to the lung parenchyma are shown. In red, vasculature staining using rhodamine-lectin. Right panel: Quantification of extravasated cells in each condition is plotted. n = 5 mice per group. 10 sections per mouse were scored. Data are averages ± SD.
C, D Adhesion to collagen I and fibronectin, respectively. The MBA-MB-231 shControl, shRARRES3 #1 and #2, LM2-Mock and LM2-RARRES3 cells were labeled with cell tracker green and plated (5 × 104) in triplicate in 24-well inserts coated with Collagen I (C) and Fibronectin (D). One hour post-plating, inserts were washed twice with PBS to remove non-attached cells and fixed in PFA. Images were taken, and the area covered by cells was determined using Image J. The percentage of area covered by cells in Collagen I (C) or Fibronectin (D) inserts is shown. Data are averages of three independent experiments ± SD (n = 3).

Homology 3D structural model of RARRES3 (red) aligned with the structure of H-REV107 (violet) (Ren et al, 2010).
The conserved catalytic residues of RARRES3 and H-REV107 showing the proposed similarity of function.
Human Arachidonic Acid (AA) levels (ng/ml) measured in cell extracts of MDA-MB-231 Parental and LM2 cellular derivatives expressing different levels of RARRES3. Data are represented as the mean of three independent experiments ± SEM.
Luciferase activity of 3x-AOX peroxisome proliferation response element reporter plasmid in MDA-MB-231 parental cells transiently transfected with Mock, RARRES3- and RARRES3-DEAD-expressing vectors. Activity of 3xAOX promoter was normalized to control condition and presented in arbitrary units. Data are mean of three independent experiments with ± SD.
The correlation coefficient and significance of RARRES3 expression levels in ER− BC primary tumors from the MSK/EMC meta-cohort against the expression of 13 PPAR target genes represented in the U133A affymetrix array are shown, together with the p value associated with each correlation. Moreover, the risk of lung metastasis (HR) associated with the expression of each of those genes in primary tumors is also reported.
Gene set enrichment analysis (GSEA) representing association of HR of lung metastasis with the PPAR target RARRES3-correlated gene set in the human breast cancer dataset (MSK/EMC expression dataset). NES-normalized enrichment score; FDR-false discovery rate; HR-hazard ratio.
The correlation coefficient of RARRES3 expression levels in ER− BC primary tumors from the MSK/EMC meta-cohort against the expression of all the genes represented in the U133A affymetrix array is shown. In red, differentiation GATA transcription factors. In black, some PPAR target genes described in (E). In green, stemness gene.

Mock- and RARRES3-expressing LM2 cells or shControl and shRARRES3 CN37 cells at the indicated dilutions were intrapulmonary injected, and grow in the lungs assessed by luciferase bioluminescence over time. Kaplan–Meier plots of the probability of lung metastasis-free survival and log-rank test were used. n = 7 mice per group was used unless indicated otherwise.
Mock- and RARRES3-expressing 4T1 cells at the indicated dilutions were intrapulmonary injected, and grow in the lungs was confirmed macroscopically 20 days post-implantation. Contingency plots and Fisher exact test were used. Representative images are shown (white dashed line limits tumor area). n = 8 mice per group were used.
Equal limiting dilutions of the indicated cell lines were plated in Matrigel. The number of organotypic spheres per plate was counted. Shown is the average of three independent experiments ± SD. Representative images are shown. Scale bar represents 50 μm.
Limiting dilutions of the indicated cell lines were performed, and one cell was plated per well of 96-well plates. One plate per cell line was cultured. The percentage of wells that generated oncospheres was calculated. Shown is the average of three independent experiments ± SD.
Human mRNA qPCR analysis of the indicated genes and cell lines in oncospheres cultures is shown. Shown is the average of at least three independent experiments ± SD.
Human mRNA qPCR or IHC analysis of the indicated genes on lung metastasis of mice inoculated with Mock and RARRES3-expressing LM2 cells is shown (n = 7). Shown is the average all samples ± SD or a representative image of these tumors.
RW.4 cells expressing a control vector (W4), a RARRES3, or a RARRES3-DEAD mutant-overexpressing vector (W4-RARRES and W4-RARRES3-DEAD) were cultured in the presence or absence of 1M all-trans retinoic acid (ATRA) for 3 days. Upper panels: representative images of the cultures are shown. Lower panels: qPCR analysis of RARRES3, GATA6, and PPIA (control) mRNAs of the indicated cultures was performed. Data are average of three independent experiments ± SD.
Comment in
-
Breast cancer: RARRES3-suppressing metastases to the lung in breast cancer.Nat Rev Clin Oncol. 2014 Jul;11(7):378. doi: 10.1038/nrclinonc.2014.103. Epub 2014 Jun 10. Nat Rev Clin Oncol. 2014. PMID: 24913373 No abstract available.
Similar articles
-
GATA3 inhibits breast cancer growth and pulmonary breast cancer metastasis.Oncogene. 2009 Jul 23;28(29):2634-42. doi: 10.1038/onc.2009.126. Epub 2009 Jun 1. Oncogene. 2009. PMID: 19483726
-
Breast cancer: RARRES3-suppressing metastases to the lung in breast cancer.Nat Rev Clin Oncol. 2014 Jul;11(7):378. doi: 10.1038/nrclinonc.2014.103. Epub 2014 Jun 10. Nat Rev Clin Oncol. 2014. PMID: 24913373 No abstract available.
-
The metastasis suppressor RARRES3 as an endogenous inhibitor of the immunoproteasome expression in breast cancer cells.Sci Rep. 2017 Jan 4;7:39873. doi: 10.1038/srep39873. Sci Rep. 2017. PMID: 28051153 Free PMC article.
-
Retinoid receptors in human lung cancer and breast cancer.Mutat Res. 1996 Feb 19;350(1):267-77. doi: 10.1016/0027-5107(95)00102-6. Mutat Res. 1996. PMID: 8657191 Review.
-
Molecular Mechanisms of Breast Cancer Metastasis to the Lung: Clinical and Experimental Perspectives.Int J Mol Sci. 2019 May 8;20(9):2272. doi: 10.3390/ijms20092272. Int J Mol Sci. 2019. PMID: 31071959 Free PMC article. Review.
Cited by
-
A Quantitative Clinicopathological Signature for Predicting Recurrence Risk of Pancreatic Ductal Adenocarcinoma After Radical Resection.Front Oncol. 2019 Nov 12;9:1197. doi: 10.3389/fonc.2019.01197. eCollection 2019. Front Oncol. 2019. PMID: 31781499 Free PMC article.
-
ATM regulation of IL-8 links oxidative stress to cancer cell migration and invasion.Elife. 2015 Jun 1;4:e07270. doi: 10.7554/eLife.07270. Elife. 2015. PMID: 26030852 Free PMC article.
-
RARRES3 suppressed metastasis through suppression of MTDH to regulate epithelial-mesenchymal transition in colorectal cancer.Am J Cancer Res. 2015 May 15;5(6):1988-99. eCollection 2015. Am J Cancer Res. 2015. PMID: 26269758 Free PMC article.
-
RARRES3 regulates signal transduction through post-translational protein modifications.Mol Cell Oncol. 2015 Feb 25;2(4):e999512. doi: 10.1080/23723556.2014.999512. eCollection 2015 Oct-Dec. Mol Cell Oncol. 2015. PMID: 27308522 Free PMC article.
-
Phospholipase A and acyltransferase 4/retinoic acid receptor responder 3 at the intersection of tumor suppression and pathogen restriction.Front Immunol. 2023 Mar 31;14:1107239. doi: 10.3389/fimmu.2023.1107239. eCollection 2023. Front Immunol. 2023. PMID: 37063830 Free PMC article. Review.
References
-
- Aslakson CJ, Miller FR. Selective events in the metastatic process defined by analysis of the sequential dissemination of subpopulations of a mouse mammary tumor. Cancer Res. 1992;52:1399–1405. - PubMed
-
- Asselin-Labat ML, Sutherland KD, Vaillant F, Gyorki DE, Wu D, Holroyd S, Breslin K, Ward T, Shi W, Bath ML, et al. Gata-3 negatively regulates the tumor-initiating capacity of mammary luminal progenitor cells and targets the putative tumor suppressor caspase-14. Mol Cell Biol. 2011;31:4609–4622. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

