Large-Scale Proteomics of the Cassava Storage Root and Identification of a Target Gene to Reduce Postharvest Deterioration
- PMID: 24876255
- PMCID: PMC4079358
- DOI: 10.1105/tpc.114.123927
Large-Scale Proteomics of the Cassava Storage Root and Identification of a Target Gene to Reduce Postharvest Deterioration
Abstract
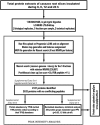
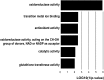
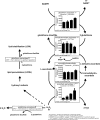
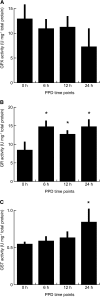
Cassava (Manihot esculenta) is the most important root crop in the tropics, but rapid postharvest physiological deterioration (PPD) of the root is a major constraint to commercial cassava production. We established a reliable method for image-based PPD symptom quantification and used label-free quantitative proteomics to generate an extensive cassava root and PPD proteome. Over 2600 unique proteins were identified in the cassava root, and nearly 300 proteins showed significant abundance regulation during PPD. We identified protein abundance modulation in pathways associated with oxidative stress, phenylpropanoid biosynthesis (including scopoletin), the glutathione cycle, fatty acid α-oxidation, folate transformation, and the sulfate reduction II pathway. Increasing protein abundances and enzymatic activities of glutathione-associated enzymes, including glutathione reductases, glutaredoxins, and glutathione S-transferases, indicated a key role for ascorbate/glutathione cycles. Based on combined proteomics data, enzymatic activities, and lipid peroxidation assays, we identified glutathione peroxidase as a candidate for reducing PPD. Transgenic cassava overexpressing a cytosolic glutathione peroxidase in storage roots showed delayed PPD and reduced lipid peroxidation as well as decreased H2O2 accumulation. Quantitative proteomics data from ethene and phenylpropanoid pathways indicate additional gene candidates to further delay PPD. Cassava root proteomics data are available at www.pep2pro.ethz.ch for easy access and comparison with other proteomics data.
© 2014 American Society of Plant Biologists. All rights reserved.
Figures

References
-
- Achidi A.U., Ajayi O.A., Bokanga M., Maziya-Dixon B. (2005). The use of cassava leaves as food in Africa. Ecol. Food Nutr. 44: 423–435
-
- Alexa A., Rahnenführer J., Lengauer T. (2006). Improved scoring of functional groups from gene expression data by decorrelating GO graph structure. Bioinformatics 22: 1600–1607 - PubMed
-
- Amako K., Chen G.X., Asada K. (1994). Separate assays specific for ascorbate peroxidase and guaiacol peroxidase and for the chloroplastic and cytosolic isozymes of ascorbate peroxidase in plants. Plant Cell Physiol. 35: 497–504
-
- Baba A.I., Nogueira F.C.S., Pinheiro C.B., Brasil J.N., Jereissati E.S., Juca T.L., Soares A.A., Santos M.F., Domont G.B., Campos F.A.P. (2008). Proteome analysis of secondary somatic embryogenesis in cassava (Manihot esculenta). Plant Sci. 175: 717–723
-
- Baerenfaller K., Hirsch-Hoffmann M., Svozil J., Hull R., Russenberger D., Bischof S., Lu Q., Gruissem W., Baginsky S. (2011). pep2pro: A new tool for comprehensive proteome data analysis to reveal information about organ-specific proteomes in Arabidopsis thaliana. Integr. Biol. (Camb.) 3: 225–237 - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

