The nucleolar size is associated to the methylation status of ribosomal DNA in breast carcinomas
- PMID: 24884608
- PMCID: PMC4062283
- DOI: 10.1186/1471-2407-14-361
The nucleolar size is associated to the methylation status of ribosomal DNA in breast carcinomas
Abstract
Background: There is a body of evidence that shows a link between tumorigenesis and ribosome biogenesis. The precursor of mature 18S, 28S and 5.8S ribosomal RNAs is transcribed from the ribosomal DNA gene (rDNA), which exists as 300-400 copies in the human diploid genome. Approximately one half of these copies are epigenetically silenced, but the exact role of epigenetic regulation on ribosome biogenesis is not completely understood. In this study we analyzed the methylation profiles of the rDNA promoter and of the 5' regions of 18S and 28S in breast cancer.
Methods: We analyzed rDNA methylation in 68 breast cancer tissues of which the normal counterpart was partially available (45/68 samples) using the MassARRAY EpiTYPER assay, a sensitive and quantitative method with single base resolution.
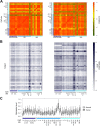
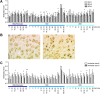
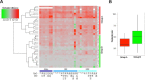
Results: We found that rDNA locus tended to be hypermethylated in tumor compared to matched normal breast tissues and that the DNA methylation level of several CpG units within the rDNA locus was associated to nuclear grade and to nucleolar size of tumor tissues. In addition we identified a subgroup of samples in which large nucleoli were associated with very limited or absent rDNA hypermethylation in tumor respect to matched normal tissue.
Conclusions: In conclusion, we suggest that rDNA is an important target of epigenetic regulation in breast tumors and that rDNA methylation level is associated to nucleolar size.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

