Analyses of clinicopathological, molecular, and prognostic associations of KRAS codon 61 and codon 146 mutations in colorectal cancer: cohort study and literature review
- PMID: 24885062
- PMCID: PMC4051153
- DOI: 10.1186/1476-4598-13-135
Analyses of clinicopathological, molecular, and prognostic associations of KRAS codon 61 and codon 146 mutations in colorectal cancer: cohort study and literature review
Abstract
Background: KRAS mutations in codons 12 and 13 are established predictive biomarkers for anti-EGFR therapy in colorectal cancer. Previous studies suggest that KRAS codon 61 and 146 mutations may also predict resistance to anti-EGFR therapy in colorectal cancer. However, clinicopathological, molecular, and prognostic features of colorectal carcinoma with KRAS codon 61 or 146 mutation remain unclear.
Methods: We utilized a molecular pathological epidemiology database of 1267 colon and rectal cancers in the Nurse's Health Study and the Health Professionals Follow-up Study. We examined KRAS mutations in codons 12, 13, 61 and 146 (assessed by pyrosequencing), in relation to clinicopathological features, and tumor molecular markers, including BRAF and PIK3CA mutations, CpG island methylator phenotype (CIMP), LINE-1 methylation, and microsatellite instability (MSI). Survival analyses were performed in 1067 BRAF-wild-type cancers to avoid confounding by BRAF mutation. Cox proportional hazards models were used to compute mortality hazard ratio, adjusting for potential confounders, including disease stage, PIK3CA mutation, CIMP, LINE-1 hypomethylation, and MSI.
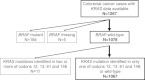
Results: KRAS codon 61 mutations were detected in 19 cases (1.5%), and codon 146 mutations in 40 cases (3.2%). Overall KRAS mutation prevalence in colorectal cancers was 40% (=505/1267). Of interest, compared to KRAS-wild-type, overall, KRAS-mutated cancers more frequently exhibited cecal location (24% vs. 12% in KRAS-wild-type; P < 0.0001), CIMP-low (49% vs. 32% in KRAS-wild-type; P < 0.0001), and PIK3CA mutations (24% vs. 11% in KRAS-wild-type; P < 0.0001). These trends were evident irrespective of mutated codon, though statistical power was limited for codon 61 mutants. Neither KRAS codon 61 nor codon 146 mutation was significantly associated with clinical outcome or prognosis in univariate or multivariate analysis [colorectal cancer-specific mortality hazard ratio (HR) = 0.81, 95% confidence interval (CI) = 0.29-2.26 for codon 61 mutation; colorectal cancer-specific mortality HR = 0.86, 95% CI = 0.42-1.78 for codon 146 mutation].
Conclusions: Tumors with KRAS mutations in codons 61 and 146 account for an appreciable proportion (approximately 5%) of colorectal cancers, and their clinicopathological and molecular features appear generally similar to KRAS codon 12 or 13 mutated cancers. To further assess clinical utility of KRAS codon 61 and 146 testing, large-scale trials are warranted.
Figures

Similar articles
-
Specific mutations in KRAS codons 12 and 13, and patient prognosis in 1075 BRAF wild-type colorectal cancers.Clin Cancer Res. 2012 Sep 1;18(17):4753-63. doi: 10.1158/1078-0432.CCR-11-3210. Epub 2012 Jul 2. Clin Cancer Res. 2012. PMID: 22753589 Free PMC article.
-
TGFBR2 and BAX mononucleotide tract mutations, microsatellite instability, and prognosis in 1072 colorectal cancers.PLoS One. 2011;6(9):e25062. doi: 10.1371/journal.pone.0025062. Epub 2011 Sep 20. PLoS One. 2011. PMID: 21949851 Free PMC article.
-
Prognostic role of PIK3CA mutation in colorectal cancer: cohort study and literature review.Clin Cancer Res. 2012 Apr 15;18(8):2257-68. doi: 10.1158/1078-0432.CCR-11-2410. Epub 2012 Feb 22. Clin Cancer Res. 2012. PMID: 22357840 Free PMC article.
-
Recommendations from the EGAPP Working Group: can testing of tumor tissue for mutations in EGFR pathway downstream effector genes in patients with metastatic colorectal cancer improve health outcomes by guiding decisions regarding anti-EGFR therapy?Genet Med. 2013 Jul;15(7):517-27. doi: 10.1038/gim.2012.184. Epub 2013 Feb 21. Genet Med. 2013. PMID: 23429431
-
Are KRAS/BRAF mutations potent prognostic and/or predictive biomarkers in colorectal cancers?Anticancer Agents Med Chem. 2012 Feb;12(2):163-71. doi: 10.2174/187152012799014968. Anticancer Agents Med Chem. 2012. PMID: 22043994 Free PMC article. Review.
Cited by
-
Alisertib exerts KRAS allele‑specific anticancer effects on colorectal cancer cell lines.Exp Ther Med. 2023 Apr 11;25(6):243. doi: 10.3892/etm.2023.11942. eCollection 2023 Jun. Exp Ther Med. 2023. PMID: 37153900 Free PMC article.
-
Molecular phenotypes of colorectal cancer and potential clinical applications.Gastroenterol Rep (Oxf). 2015 Nov;3(4):269-76. doi: 10.1093/gastro/gov046. Epub 2015 Sep 3. Gastroenterol Rep (Oxf). 2015. PMID: 26337942 Free PMC article. Review.
-
Functional Characterization and Drug Response of Freshly Established Patient-Derived Tumor Models with CpG Island Methylator Phenotype.PLoS One. 2015 Nov 30;10(11):e0143194. doi: 10.1371/journal.pone.0143194. eCollection 2015. PLoS One. 2015. PMID: 26618628 Free PMC article.
-
Specific mutations in KRAS codon 12 are associated with worse overall survival in patients with advanced and recurrent colorectal cancer.Br J Cancer. 2017 Mar 28;116(7):923-929. doi: 10.1038/bjc.2017.37. Epub 2017 Feb 16. Br J Cancer. 2017. PMID: 28208157 Free PMC article.
-
Aspirin exerts high anti-cancer activity in PIK3CA-mutant colon cancer cells.Oncotarget. 2017 Sep 18;8(50):87379-87389. doi: 10.18632/oncotarget.20972. eCollection 2017 Oct 20. Oncotarget. 2017. PMID: 29152088 Free PMC article.
References
-
- Febbo PG, Ladanyi M, Aldape KD, De Marzo AM, Hammond ME, Hayes DF, Iafrate AJ, Kelley RK, Marcucci G, Ogino S, Pao W, Sgroi DC, Birkeland ML. NCCN Task Force report: evaluating the clinical utility of tumor markers in oncology. J Natl Compr Canc Netw. 2011;9(Suppl 5):S1–S32. - PubMed
-
- Rosty C, Young JP, Walsh MD, Clendenning M, Walters RJ, Pearson S, Pavluk E, Nagler B, Pakenas D, Jass JR, Jenkins MA, Win AK, Southey MC, Parry S, Hopper JL, Giles GG, Williamson E, English DR, Buchanan DD. Colorectal carcinomas with KRAS mutation are associated with distinctive morphological and molecular features. Mod Pathol. 2013;26:825–834. doi: 10.1038/modpathol.2012.240. - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

