Alcohol dehydrogenases from Kluyveromyces marxianus: heterologous expression in Escherichia coli and biochemical characterization
- PMID: 24885162
- PMCID: PMC4062290
- DOI: 10.1186/1472-6750-14-45
Alcohol dehydrogenases from Kluyveromyces marxianus: heterologous expression in Escherichia coli and biochemical characterization
Abstract
Background: Kluyveromyces marxianus has recently become a species of interest for ethanol production since it can produce ethanol at high temperature and on a wide variety of substrates. However, the reason why this yeast can produce ethanol at high temperature is largely unknown.
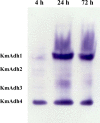
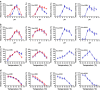
Results: The ethanol fermentation capability of K. marxianus GX-UN120 at 40°С was found to be the same as that of Saccharomyces cerevisiae at 34°С. Zymogram analysis showed that alcohol dehydrogenase 1 (KmAdh1) was largely induced during ethanol production, KmAdh4 was constitutively expressed at a lower level and KmAdh2 and KmAdh3 were almost undetectable. The genes encoding the four alcohol dehydrogenases (ADHs) were cloned from strain GX-UN120. Each KmADH was expressed in Escherichia coli and each recombinant protein was digested with enterokinase to remove the fusion protein. The optimum pH of the purified recombinant KmAdh1 was 8.0 and that of KmAdh2, KmAdh3 and KmAdh4 was 7.0. The optimum temperatures of KmAdh1, KmAdh2, KmAdh3 and KmAdh4 were 50, 45, 55 and 45°C, respectively. The K(m) values of the recombinant KmAdh1 and KmAdh2 were 4.0 and 1.2 mM for acetaldehyde and 39.7 and 49.5 mM for ethanol, respectively. The V(max) values of the recombinant KmAdh1 and KmAdh2 were 114.9 and 21.6 μmol min⁻¹ mg⁻¹ for acetaldehyde and 57.5 and 1.8 μmol min⁻¹ mg⁻¹ for ethanol, respectively. KmAdh3 and KmAdh4 catalyze the oxidation reaction of ethanol to acetaldehyde but not the reduction reaction of acetaldehyde to ethanol, and the K(m) values of the recombinant KmAdh3 and KmAdh4 were 26.0 and 17.0 mM for ethanol, respectively. The V(max) values of the recombinant KmAdh3 and KmAdh4 were 12.8 and 56.2 μmol min⁻¹ mg⁻¹ for ethanol, respectively.
Conclusion: These data in this study collectively indicate that KmAdh1 is the primary ADH responsible for the production of ethanol from the reduction of acetaldehyde in K. marxianus. The relatively high optimum temperature of KmAdh1 may partially explain the ability of K. marxianus to produce ethanol at high temperature. Understanding the biochemical characteristics of KmAdhs will enhance our fundamental knowledge of the metabolism of ethanol fermentation in K. marxianus.
Figures





Similar articles
-
Comparison of the gene expression patterns of alcohol dehydrogenase isozymes in the thermotolerant yeast Kluyveromyces marxianus and their physiological functions.Biosci Biotechnol Biochem. 2007 May;71(5):1170-82. doi: 10.1271/bbb.60622. Epub 2007 May 7. Biosci Biotechnol Biochem. 2007. PMID: 17485854
-
Identification of a xylitol dehydrogenase gene from Kluyveromyces marxianus NBRC1777.Mol Biotechnol. 2013 Feb;53(2):159-69. doi: 10.1007/s12033-012-9508-9. Mol Biotechnol. 2013. PMID: 22351371
-
High-temperature ethanol production by a series of recombinant xylose-fermenting Kluyveromyces marxianus strains.Enzyme Microb Technol. 2019 Oct;129:109359. doi: 10.1016/j.enzmictec.2019.109359. Epub 2019 Jun 14. Enzyme Microb Technol. 2019. PMID: 31307575
-
Kluyveromyces marxianus as a host for heterologous protein synthesis.Appl Microbiol Biotechnol. 2016 Jul;100(14):6193-6208. doi: 10.1007/s00253-016-7645-y. Epub 2016 Jun 3. Appl Microbiol Biotechnol. 2016. PMID: 27260286 Review.
-
The yeast Kluyveromyces marxianus and its biotechnological potential.Appl Microbiol Biotechnol. 2008 Jun;79(3):339-54. doi: 10.1007/s00253-008-1458-6. Epub 2008 Apr 22. Appl Microbiol Biotechnol. 2008. PMID: 18427804 Review.
Cited by
-
Effects of glucose, ethanol and acetic acid on regulation of ADH2 gene from Lachancea fermentati.PeerJ. 2016 Mar 10;4:e1751. doi: 10.7717/peerj.1751. eCollection 2016. PeerJ. 2016. PMID: 26989608 Free PMC article.
-
The novel properties of Kluyveromyces marxianus glucose sensor/receptor repressor pathway and the construction of glucose repression-released strains.Microb Cell Fact. 2023 Jul 10;22(1):123. doi: 10.1186/s12934-023-02138-7. Microb Cell Fact. 2023. PMID: 37430283 Free PMC article.
-
Extensive remodeling of sugar metabolism through gene loss and horizontal gene transfer in a eukaryotic lineage.BMC Biol. 2024 May 30;22(1):128. doi: 10.1186/s12915-024-01929-7. BMC Biol. 2024. PMID: 38816863 Free PMC article.
-
Establishment of Kluyveromyces marxianus as a Microbial Cell Factory for Lignocellulosic Processes: Production of High Value Furan Derivatives.J Fungi (Basel). 2021 Dec 7;7(12):1047. doi: 10.3390/jof7121047. J Fungi (Basel). 2021. PMID: 34947029 Free PMC article.
-
Characterization of argonaute nucleases from mesophilic bacteria Pseudobutyrivibrio ruminis.Bioresour Bioprocess. 2024 Oct 7;11(1):94. doi: 10.1186/s40643-024-00797-x. Bioresour Bioprocess. 2024. PMID: 39373873 Free PMC article.
References
-
- Lane MM, Morrissey JP. Kluyveromyces marxianus: a yeast emerging from its sister’s shadow. Fungal Biol Rev. 2010;24:17–26. doi: 10.1016/j.fbr.2010.01.001. - DOI
-
- Nonklang S, Abdel-Banat BMA, Cha-aim K, Moonjai N, Hoshida H, Limtong S, Yamada M, Akada R. High-temperature ethanol fermentation and transformation with linear DNA in the thermotolerant yeast Kluyveromyces marxianus DMKU3-1042. Appl Environ Microbiol. 2008;74:7514–7521. doi: 10.1128/AEM.01854-08. - DOI - PMC - PubMed
-
- Wilkins MR, Mueller M, Eichling S, Banat IM. Fermentation of xylose by the thermotolerant yeast strains Kluyveromyces marxianus IMB2, IMB4, and IMB5 under anaerobic conditions. Process Biochem. 2008;43:346–350. doi: 10.1016/j.procbio.2007.12.011. - DOI
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

