Cloning, expression and biochemical characterization of the cholesterol oxidase CgChoA from Chryseobacterium gleum
- PMID: 24885249
- PMCID: PMC4053396
- DOI: 10.1186/1472-6750-14-46
Cloning, expression and biochemical characterization of the cholesterol oxidase CgChoA from Chryseobacterium gleum
Abstract
Background: Cholesterol oxidases are important enzymes for applications such as the analysis of cholesterol in clinical samples, the synthesis of steroid derived drugs, and are considered as potential antibacterial drug targets.
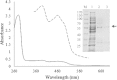
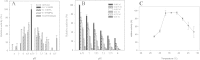
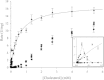
Results: The gene choA encoding a cholesterol oxidase from Chryseobacterium gleum DSM 16776 was cloned into the pQE-30 expression vector and heterologously expressed in Escherichia coli JM109 co-transformed with pRARE2. The N-terminally His-tagged cholesterol oxidase (CgChoA) was assigned to be a monomer in solution by size exclusion chromatography, showed a temperature optimum of 35°C, and a pH optimum at 6.75 using 0.011 M MOPS buffer under the tested conditions. The purified protein showed a maximum activity of 15.5 U/mg. CgChoA showed a Michaelis-Menten like kinetic behavior only when the substrate was dissolved in water and taurocholate (apparent K(m) = 0.5 mM). In addition, the conversion of cholesterol by CgChoA was studied via biocatalytic batches at analytical scale, and cholest-4-en-3-one was confirmed as product by HPLC-MS.
Conclusion: CgChoA is a true cholesterol oxidase which activity ranges among the high performing described cholesterol oxidases from other organisms. Thus, the enzyme broadens the available toolbox of cholesterol oxidases for e.g. synthetic and biosensing applications.
Figures
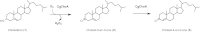


Similar articles
-
Cholesterol oxidase with high catalytic activity from Pseudomonas aeruginosa: Screening, molecular genetic analysis, expression and characterization.J Biosci Bioeng. 2015 Jul;120(1):24-30. doi: 10.1016/j.jbiosc.2014.12.003. Epub 2015 Jan 5. J Biosci Bioeng. 2015. PMID: 25573142
-
Purification and properties of a new Brevibacterium sterolicum cholesterol oxidase produced by E. coli MM294/pnH10.FEMS Microbiol Lett. 2002 Oct 8;215(2):243-8. doi: 10.1111/j.1574-6968.2002.tb11397.x. FEMS Microbiol Lett. 2002. PMID: 12399041
-
Cholesterol biotransformation to androsta-1,4-diene-3,17-dione by growing cells of Chryseobacterium gleum.Biotechnol Lett. 2010 May;32(5):695-9. doi: 10.1007/s10529-010-0206-z. Epub 2010 Feb 3. Biotechnol Lett. 2010. PMID: 20127502
-
Cholesterol oxidase: biotechnological applications.FEBS J. 2009 Dec;276(23):6857-70. doi: 10.1111/j.1742-4658.2009.07379.x. Epub 2009 Oct 16. FEBS J. 2009. PMID: 19843167 Review.
-
Characteristics and biotechnological applications of microbial cholesterol oxidases.Appl Microbiol Biotechnol. 2009 Jul;83(5):825-37. doi: 10.1007/s00253-009-2059-8. Epub 2009 Jun 3. Appl Microbiol Biotechnol. 2009. PMID: 19495743 Review.
Cited by
-
Cloning, expression, and in silico structural modeling of cholesterol oxidase of Acinetobacter sp. strain RAMD in E. coli.FEBS Open Bio. 2021 Sep;11(9):2560-2575. doi: 10.1002/2211-5463.13254. Epub 2021 Jul 31. FEBS Open Bio. 2021. PMID: 34272838 Free PMC article.
-
Expression optimization, purification, and functional characterization of cholesterol oxidase from Chromobacterium sp. DS1.PLoS One. 2019 Feb 13;14(2):e0212217. doi: 10.1371/journal.pone.0212217. eCollection 2019. PLoS One. 2019. PMID: 30759160 Free PMC article.
-
Characterization and overproduction of cell-associated cholesterol oxidase ChoD from Streptomyces lavendulae YAKB-15.Sci Rep. 2019 Aug 14;9(1):11850. doi: 10.1038/s41598-019-48132-1. Sci Rep. 2019. PMID: 31413341 Free PMC article.
-
Microbial transformation of cholesterol: reactions and practical aspects-an update.World J Microbiol Biotechnol. 2019 Aug 20;35(9):131. doi: 10.1007/s11274-019-2708-8. World J Microbiol Biotechnol. 2019. PMID: 31432251 Review.
References
-
- Kumari L, Kanwar SS. Cholesterol Oxidase and Its Applications. Advances in Microbiology. 2012;2(2):49–65. doi: 10.4236/aim.2012.22007. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

