Nucleomorph and plastid genome sequences of the chlorarachniophyte Lotharella oceanica: convergent reductive evolution and frequent recombination in nucleomorph-bearing algae
- PMID: 24885563
- PMCID: PMC4035089
- DOI: 10.1186/1471-2164-15-374
Nucleomorph and plastid genome sequences of the chlorarachniophyte Lotharella oceanica: convergent reductive evolution and frequent recombination in nucleomorph-bearing algae
Abstract
Background: Nucleomorphs are residual nuclei derived from eukaryotic endosymbionts in chlorarachniophyte and cryptophyte algae. The endosymbionts that gave rise to nucleomorphs and plastids in these two algal groups were green and red algae, respectively. Despite their independent origin, the chlorarachniophyte and cryptophyte nucleomorph genomes share similar genomic features such as extreme size reduction and a three-chromosome architecture. This suggests that similar reductive evolutionary forces have acted to shape the nucleomorph genomes in the two groups. Thus far, however, only a single chlorarachniophyte nucleomorph and plastid genome has been sequenced, making broad evolutionary inferences within the chlorarachniophytes and between chlorarachniophytes and cryptophytes difficult. We have sequenced the nucleomorph and plastid genomes of the chlorarachniophyte Lotharella oceanica in order to gain insight into nucleomorph and plastid genome diversity and evolution.
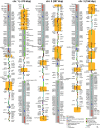
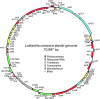
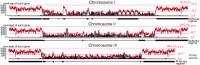
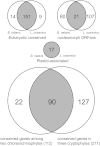
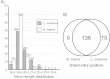
Results: The L. oceanica nucleomorph genome was found to consist of three linear chromosomes totaling ~610 kilobase pairs (kbp), much larger than the 373 kbp nucleomorph genome of the model chlorarachniophyte Bigelowiella natans. The L. oceanica plastid genome is 71 kbp in size, similar to that of B. natans. Unexpectedly long (~35 kbp) sub-telomeric repeat regions were identified in the L. oceanica nucleomorph genome; internal multi-copy regions were also detected. Gene content analyses revealed that nucleomorph house-keeping genes and spliceosomal intron positions are well conserved between the L. oceanica and B. natans nucleomorph genomes. More broadly, gene retention patterns were found to be similar between nucleomorph genomes in chlorarachniophytes and cryptophytes. Chlorarachniophyte plastid genomes showed near identical protein coding gene complements as well as a high level of synteny.
Conclusions: We have provided insight into the process of nucleomorph genome evolution by elucidating the fine-scale dynamics of sub-telomeric repeat regions. Homologous recombination at the chromosome ends appears to be frequent, serving to expand and contract nucleomorph genome size. The main factor influencing nucleomorph genome size variation between different chlorarachniophyte species appears to be expansion-contraction of these telomere-associated repeats rather than changes in the number of unique protein coding genes. The dynamic nature of chlorarachniophyte nucleomorph genomes lies in stark contrast to their plastid genomes, which appear to be highly stable in terms of gene content and synteny.
Figures

References
Publication types
MeSH terms
Associated data
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

