Porcine circovirus type 2 in China: an update on and insights to its prevalence and control
- PMID: 24885983
- PMCID: PMC4031328
- DOI: 10.1186/1743-422X-11-88
Porcine circovirus type 2 in China: an update on and insights to its prevalence and control
Abstract
Currently, porcine circovirus type 2 (PCV2) is considered the major pathogen of porcine circovirus associated-diseases (PCVAD) that causes large economic losses for the swine industry in the world annually, including China. Since the first report of PCV2 in 1998, it has been drawing tremendous attention for the government, farming enterprises, farmers, and veterinary practitioners. Chinese researchers have conducted a number of molecular epidemiological work on PCV2 by molecular approaches in the past several years, which has resulted in the identification of novel PCV2 genotypes and PCV2-like agents as well as the description of new prevalence patterns. Since late 2009, commercial PCV2 vaccines, including the subunit vaccines and inactivated vaccines, have already been used in Chinese swine farms. The aim of this review is to update the insights into the prevalence and control of PCV2 in China, which would contribute to understanding the epidemiology, control measures and design of novel vaccines for PCV2.
Figures
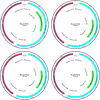
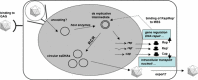
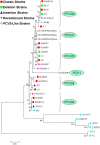

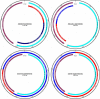
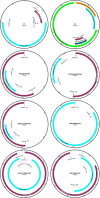
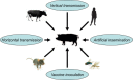
References
-
- Tischer I, Rasch R, Tochtermann G. Characterization of papovavirus-and picornavirus-like particles in permanent pig kidney cell lines. Zentralbl Bakteriol Orig A. 1974;226:153–167. - PubMed
-
- Mankertz A, Mankertz J, Wolf K, Buhk HJ. Identification of a protein essential for replication of porcine circovirus. J Gen Virol. 1998;79:381–384. - PubMed
-
- Nawagitgul P, Morozov I, Bolin SR, Harms PA, Sorden SD, Paul PS. Open reading frame 2 of porcine circovirus type 2 encodes a major capsid protein. J Gen Virol. 2000;81:2281–2287. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

